 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG001955t1 | ||||||||
| Common Name | TCM_001955 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 290aa MW: 33283.6 Da PI: 7.7402 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 161.1 | 4.3e-50 | 18 | 146 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevl 93
l pGfrFhPtdeelv +yL++kve++++++ e+ik++diy+++PwdLp+ + ++kewyfF+ r +ky+++ r+nr+t sg+Wkatg dk+++
Thecc1EG001955t1 18 LLPGFRFHPTDEELVGFYLRRKVENRPISI-EIIKQIDIYRYDPWDLPRVSTVGDKEWYFFCIRGRKYRNSIRPNRVTGSGFWKATGIDKPIY 109
579***************************.99***************7777799************************************** PP
NAM 94 sk..kgelvglkktLvfykgrapkgektdWvmheyrl 128
+ + +glkk+Lv+y+g+a kg+ktdW+mhe+rl
Thecc1EG001955t1 110 TVkePHDCIGLKKSLVYYRGSAGKGTKTDWMMHEFRL 146
97434445***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.01E-56 | 9 | 167 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 55.246 | 18 | 168 | IPR003441 | NAC domain |
| Pfam | PF02365 | 2.8E-25 | 20 | 146 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 290 aa Download sequence Send to blast |
MEATNMNSTD KGKDDEDLLP GFRFHPTDEE LVGFYLRRKV ENRPISIEII KQIDIYRYDP 60 WDLPRVSTVG DKEWYFFCIR GRKYRNSIRP NRVTGSGFWK ATGIDKPIYT VKEPHDCIGL 120 KKSLVYYRGS AGKGTKTDWM MHEFRLPPNA KDVAQEAEVW TLCRIFKRIP SNKKDAAAAW 180 KDTRNKQNGA YHTSSRTCSL DSENSEQCKS FCDSVLLQND IKPNIMDQVD ARNYFLVGPW 240 TNAMSQQAPF AASCPSFWNP NGDVDIFTNG NWDELRPLVE LALEPSPYL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 9e-51 | 18 | 174 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 9e-51 | 18 | 174 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 9e-51 | 18 | 174 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 9e-51 | 18 | 174 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 9e-51 | 18 | 174 | 17 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 9e-51 | 18 | 174 | 17 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
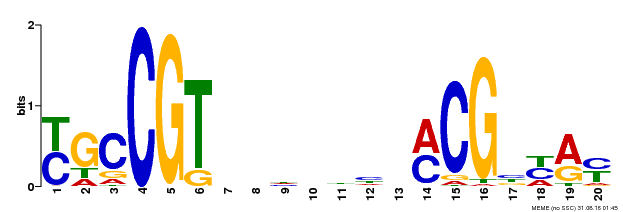
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007048954.1 | 0.0 | PREDICTED: transcription factor JUNGBRUNNEN 1 | ||||
| Swissprot | Q9SK55 | 1e-94 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | A0A061DM21 | 0.0 | A0A061DM21_THECC; NAC domain-containing protein 42 | ||||
| STRING | EOX93111 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM306 | 28 | 200 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43000.1 | 5e-97 | NAC domain containing protein 42 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG001955t1 |
| Entrez Gene | 18612219 |



