 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG000780t1 | ||||||||
| Common Name | TCM_000780 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 234aa MW: 25045.8 Da PI: 7.2753 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.2 | 0.00026 | 78 | 100 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
ykC+ C+k F++ L H +H
Thecc1EG000780t1 78 YKCTVCNKAFPSYQALGGHKASH 100
9***********99999998887 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 7.05E-10 | 77 | 100 | No hit | No description |
| Pfam | PF13912 | 7.2E-14 | 78 | 102 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0062 | 78 | 100 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.889 | 78 | 100 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 80 | 100 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.05E-10 | 138 | 165 | No hit | No description |
| Pfam | PF13912 | 9.8E-11 | 142 | 167 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.265 | 143 | 165 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.74 | 143 | 165 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 145 | 165 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 234 aa Download sequence Send to blast |
MALEMLNSPT SAPPLLHHDD IDLHSTEPWT KRKRTKRPRI ENPPTEEEYL ALCLLMLAQG 60 TTTRTSPSAA AKNTLNNYKC TVCNKAFPSY QALGGHKASH RKLGGADEHP TTTTTAAAVS 120 APTTTTATVA TNSPPMNQGG KTHTCSICFK TFSSGQALGG HKRCHYEAGS NINSNSGEGV 180 KLSSQSQRDF DLNWPAGPED PSIDVDQKDK LSGDEEEVES PSPPAKRGSY HVC* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor involved in the inhibition of plant growth under abiotic stress conditions. Can repress the expression of various genes, including osmotic stress and abscisic acid-repressive genes and auxin-inducible genes, by binding to their promoter regions in a DNA sequence-specific manner. {ECO:0000269|PubMed:15333755, ECO:0000269|PubMed:21852415}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00118 | ampDAP | Transfer from AT5G67450 | Download |
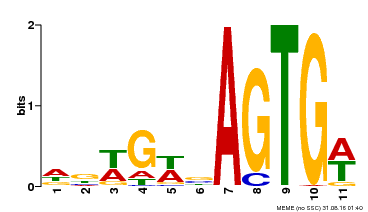
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA), ethylene, dehydration, salt and cold. {ECO:0000269|PubMed:10806347, ECO:0000269|PubMed:15333755, ECO:0000269|PubMed:21852415}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007047505.2 | 1e-171 | PREDICTED: zinc finger protein ZAT10 | ||||
| Swissprot | Q9SSW1 | 3e-48 | AZF1_ARATH; Zinc finger protein AZF1 | ||||
| TrEMBL | A0A061DHM6 | 1e-173 | A0A061DHM6_THECC; Zinc-finger DNA binding protein, putative | ||||
| STRING | EOX91662 | 1e-174 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3121 | 22 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G27730.1 | 2e-42 | salt tolerance zinc finger | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG000780t1 |
| Entrez Gene | 18611274 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



