 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG000649t3 | ||||||||
| Common Name | TCM_000649 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 390aa MW: 42129.8 Da PI: 4.856 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 49.5 | 7.4e-16 | 146 | 192 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn E+rRR+riN+++ L+ l+P++ K +Ka++L +A+eY+k+Lq
Thecc1EG000649t3 146 VHNLSEKRRRSRINEKMKALQNLIPNS-----NKTDKASMLDEAIEYLKQLQ 192
6*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 2.88E-20 | 139 | 202 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.6E-20 | 139 | 202 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.86 | 142 | 191 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.39E-17 | 145 | 196 | No hit | No description |
| Pfam | PF00010 | 2.5E-13 | 146 | 192 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.3E-18 | 148 | 197 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010114 | Biological Process | response to red light | ||||
| GO:0010154 | Biological Process | fruit development | ||||
| GO:0010187 | Biological Process | negative regulation of seed germination | ||||
| GO:0048440 | Biological Process | carpel development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 390 aa Download sequence Send to blast |
MGDNINVNMF HYNNNYTTTN DNNSNNGKCL SSASDDISNL LHQILVHSSS PSSGMAYLEG 60 PTENPRRLSR SPAPAGGEAK QGMILTVDSC GRGSGGSGLV GVAGEINDAE EYDCESEEGL 120 EALVDEAPSK PAPPRSSSKR SRAAEVHNLS EKRRRSRINE KMKALQNLIP NSNKTDKASM 180 LDEAIEYLKQ LQLQVQMLTM RNGLSLHPMC LPGVLQPIQL PQTRIDFGED NGSLPMNASG 240 TAPANQEPSA QIVFDLPNQC SSSNHALVPN MSNIITSETS FSLESIQAPF GPFQLLTPTQ 300 DICREDILPH HQLKSNTSEF GSGATSTVSL PFDTRESDLK ESSSLDASMK GRDQPNSVLE 360 HDLVLAPHLT RQAGRSDSSD DIKIEKPNF* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that plays a role in floral organogenesis. Promotes the growth of carpel margins and of pollen tract tissues derived from them. {ECO:0000269|PubMed:10225997, ECO:0000269|Ref.8}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00035 | PBM | Transfer from AT4G36930 | Download |
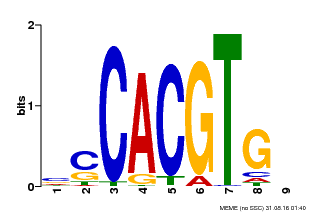
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by the A class gene AP2 in the first whorl and by ARF3/ETT in gynoecium. {ECO:0000269|PubMed:11245574}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007047306.2 | 0.0 | PREDICTED: transcription factor SPATULA isoform X2 | ||||
| Swissprot | Q9FUA4 | 2e-56 | SPT_ARATH; Transcription factor SPATULA | ||||
| TrEMBL | A0A061DNA7 | 0.0 | A0A061DNA7_THECC; Homeodomain-like superfamily protein, putative isoform 3 | ||||
| STRING | EOX91461 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36930.1 | 2e-55 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG000649t3 |
| Entrez Gene | 18611155 |



