 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_5BL_3503725F1.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 966aa MW: 105557 Da PI: 5.571 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128 | 3.7e-40 | 153 | 230 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC adls ak+yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Traes_5BL_3503725F1.2 153 ACQVEGCCADLSAAKDYHRRHKVCEMHAKANTAVVGNTIQRFCQQCSRFHLLQEFDEGKRSCRRRLAGHNKRRRKTRP 230
5**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.0E-33 | 146 | 215 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.308 | 151 | 228 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.75E-37 | 152 | 231 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 3.6E-29 | 154 | 227 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS50297 | 8.667 | 745 | 812 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 2.5 | 745 | 774 | IPR002110 | Ankyrin repeat |
| Gene3D | G3DSA:1.25.40.20 | 3.8E-7 | 747 | 853 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 8.08E-8 | 747 | 852 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.20E-8 | 750 | 850 | No hit | No description |
| SMART | SM00248 | 380 | 791 | 823 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 966 aa Download sequence Send to blast |
MEAARVGSQS RHLYGGGLGG VLEQARREKR VFGWDLNDWS WDSERFVATP VPAAVGNGLS 60 LNSTPSSSEE AEAEVARNGN GRVDSDKRKR VVVIHDDEDE KDEDPVGNSN NVLSLRIGGS 120 SVAGGTAEDG GVNEEDRNGK KIRVQGGSSS GPACQVEGCC ADLSAAKDYH RRHKVCEMHA 180 KANTAVVGNT IQRFCQQCSR FHLLQEFDEG KRSCRRRLAG HNKRRRKTRP ENAVGGTPIE 240 EKVSSYLLLS LLGICANLNS DNAEHLQGQE LLSNLWRNLG NVAKSLDPKE LCKLLEACQS 300 MQNRSNAGTS EAANALVNTA AAEAAGPSNS KAPFANGGDC GQTSSAVLPV QSNATMVATT 360 GSVFTETPAC KFRNFDLNDT CNDMEGFEDG SNCPSWIRQD STQSPPQTSG NSDSTSAQSL 420 SSSNGDAQCR TDKIVFKLFE KVPSELPPIL RSQILGWLSS SPTDIESHIR PGCIILTVYL 480 RLVESSWREL SENMSVYLDK LSSSSADNFW TSGLVFVMVR HQIAFMHNGQ VMLDRPLAPN 540 SHHYCKVLCV SPVAAPYSAT VNFRVEGFNL VSSSSRLICS IEGRCIFEED TSIMADDTED 600 EDIEYLNFCC SLPGTRGRGF IEVEDSGFSN GFFPFIVAEQ NVCSEVCELE SIFKSSSLEQ 660 PDNDIAMNQA LEFLHELGWL LHRVNIISKH DKVEPPVPAF NLLRFRNLGI FAMEREWCAV 720 TKMLLDLLFD GFVDAGLQSP KDVVLSENLL HSAVRGKSAR MVRFLLTYKP NKNLKETAET 780 YLFRPDAQGP SAFTPLHIAA ATSDAEDVLD ALTDDPGLVG LNAWRNARDE IGFTPEDYAR 840 QSGNDAYINL VQKKIDKHLG KGHVVLGVPS SMCPGITDGM KAGDISLEIC KAMPMTTTSA 900 ARCNICSCQG KMYPNSLART FLYRPAMFTV MGVAVICVCV GILLHTLPKV YAAPNFRWEL 960 LERGAM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 9e-31 | 147 | 227 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
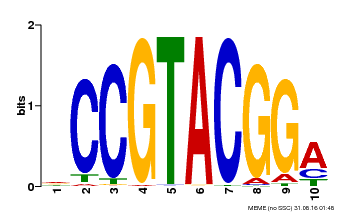
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF447880 | 0.0 | KF447880.1 Triticum aestivum squamosa promoter-binding-like protein 6 (SPL6) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020169994.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A3B6LXG5 | 0.0 | A0A3B6LXG5_WHEAT; Uncharacterized protein | ||||
| STRING | Traes_5BL_3503725F1.2 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



