 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_5BL_06A206038.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 330aa MW: 33924.4 Da PI: 8.7474 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 221.3 | 2e-68 | 97 | 248 | 3 | 154 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasaaeaaskrkrelkskkqsalsstk 90
g++sCqdCGnqakkdC+h+RCRtCCksrgf C+thvkstWvpa+krrerqqql+a++++aaa++ a + r+ +++ +++ a+++ +
Traes_5BL_06A206038.1 97 GGGTSCQDCGNQAKKDCQHQRCRTCCKSRGFACSTHVKSTWVPASKRRERQQQLTALAASAAATTGGAGPSRDLTKRPRARLAATTPT 184
5778*********************************************************999988888888888899999999999 PP
DUF702 91 lssaeskkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGlee 154
+ss+++++ + ++++P+evsseavfrcvr++ vd++e+elaYqt+vsigGhvfkGiL+d G+++
Traes_5BL_06A206038.1 185 TSSGDQQMVTVAERFPREVSSEAVFRCVRLGPVDQAEAELAYQTTVSIGGHVFKGILHDVGPSN 248
*************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 2.2E-64 | 99 | 247 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 2.2E-26 | 101 | 143 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 3.4E-23 | 198 | 246 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009938 | Biological Process | negative regulation of gibberellic acid mediated signaling pathway | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010252 | Biological Process | auxin homeostasis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048479 | Biological Process | style development | ||||
| GO:0048480 | Biological Process | stigma development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 330 aa Download sequence Send to blast |
MAGFPLGTGG SGRDSSASPT GVHPSDASTF LYATRGGGFQ LWGQPQDQQQ LTHPFYAPNL 60 IRFATDDIPA GAAQSLAGAA SSSSSRGARA AALAGTGGGT SCQDCGNQAK KDCQHQRCRT 120 CCKSRGFACS THVKSTWVPA SKRRERQQQL TALAASAAAT TGGAGPSRDL TKRPRARLAA 180 TTPTTSSGDQ QMVTVAERFP REVSSEAVFR CVRLGPVDQA EAELAYQTTV SIGGHVFKGI 240 LHDVGPSNAH AQLQAAAGGS GGGGGDYQFR LTGDVSPPST GAEAGDAGGG NNHNIVVSSA 300 VVMDPYPTPG LYGSFPATTP FFHGHPYQRQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). {ECO:0000269|PubMed:12361963, ECO:0000269|PubMed:16740145, ECO:0000269|PubMed:16740146, ECO:0000269|PubMed:18811619, ECO:0000269|PubMed:20154152, ECO:0000269|PubMed:22318676}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00613 | PBM | Transfer from AT3G51060 | Download |
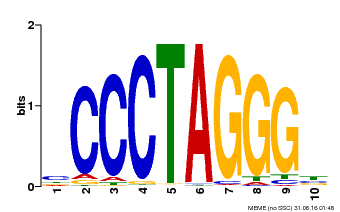
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Regulated by ESR1 and ESR2. {ECO:0000269|PubMed:21976484}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK241285 | 1e-161 | AK241285.1 Oryza sativa Japonica Group cDNA, clone: J065137A19, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020192267.1 | 0.0 | protein SHI RELATED SEQUENCE 1-like | ||||
| Swissprot | Q9SD40 | 2e-49 | SRS1_ARATH; Protein SHI RELATED SEQUENCE 1 | ||||
| TrEMBL | A0A3B6LQX1 | 0.0 | A0A3B6LQX1_WHEAT; Uncharacterized protein | ||||
| TrEMBL | A0A446UH72 | 0.0 | A0A446UH72_TRITD; Uncharacterized protein | ||||
| STRING | Traes_5BL_06A206038.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1354 | 35 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G51060.1 | 4e-43 | Lateral root primordium (LRP) protein-related | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



