 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_4DL_A2012EA68.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 260aa MW: 26929.5 Da PI: 7.8516 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 129 | 2.2e-40 | 39 | 139 | 1 | 100 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeq.pkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvilklqqql 87
+C aCk+lrrkC+++C++apyf +eq +++fa+vhk+FGasnv+kll ++p ++r da+ +++yeA+ar+rdPvyG+v++i++lqqq+
Traes_4DL_A2012EA68.1 39 PCGACKFLRRKCVSECIFAPYFDSEQgAAHFAAVHKVFGASNVSKLLLQIPAHKRLDAVVTICYEAQARLRDPVYGCVAHIFALQQQV 126
7***********************9989************************************************************ PP
DUF260 88 eqlkaelallkee 100
+l+ael +l+++
Traes_4DL_A2012EA68.1 127 VNLQAELTYLQTH 139
*******999876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50891 | 23.084 | 38 | 140 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 1.5E-39 | 39 | 137 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010311 | Biological Process | lateral root formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 260 aa Download sequence Send to blast |
MSSGGGSSTL GGCGGPSGSG SGSGGGGGGG GLGGGGGGPC GACKFLRRKC VSECIFAPYF 60 DSEQGAAHFA AVHKVFGASN VSKLLLQIPA HKRLDAVVTI CYEAQARLRD PVYGCVAHIF 120 ALQQQVVNLQ AELTYLQTHL ATLELPSPPL PAAPQMPMAM PAQFSISDLP STTNIPTTID 180 LSALFEPPAQ PQWALQQHHQ HQLRQQPSYG AMAHRGGSSS MAEGSVGSGD LQTLARELLD 240 RHGRSGVKPE LQPPPPPHPR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 3e-37 | 35 | 140 | 7 | 111 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 3e-37 | 35 | 140 | 7 | 111 | LOB family transfactor Ramosa2.1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the positive regulation of tracheary element (TE) differentiation. Involved in a positive feedback loop that maintains or promotes NAC030/VND7 expression that regulates TE differentiation-related genes (PubMed:19088331). Functions in the initiation and emergence of lateral roots, in conjunction with LBD16, downstream of ARF7 and ARF19 (PubMed:19717544, PubMed:23749813). Transcriptional activator that directly regulates EXPA14, a gene encoding a cell wall-loosening factor that promotes lateral root emergence. Activates EXPA14 by directly binding to a specific region of its promoter (PubMed:22974309). Transcriptional activator that directly regulates EXPA17, a gene encoding a cell wall-loosening factor that promotes lateral root emergence (PubMed:23872272). Acts downstream of the auxin influx carriers AUX1 and LAX1 in the regulation of lateral root initiation and development (PubMed:26059335). {ECO:0000269|PubMed:19088331, ECO:0000269|PubMed:19717544, ECO:0000269|PubMed:22974309, ECO:0000269|PubMed:23749813, ECO:0000269|PubMed:23872272, ECO:0000269|PubMed:26059335}. | |||||
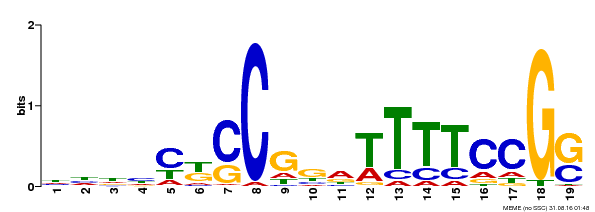
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00314 | DAP | Transfer from AT2G45420 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:15659631, ECO:0000269|PubMed:19717544, ECO:0000269|PubMed:23749813}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP094077 | 0.0 | FP094077.1 Phyllostachys edulis cDNA clone: bphylf043i04, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020160420.1 | 0.0 | LOB domain-containing protein 30 | ||||
| Swissprot | O22131 | 1e-68 | LBD18_ARATH; LOB domain-containing protein 18 | ||||
| TrEMBL | A0A3B6JKZ1 | 0.0 | A0A3B6JKZ1_WHEAT; Uncharacterized protein | ||||
| STRING | Traes_4DL_A2012EA68.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3346 | 37 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45420.1 | 1e-63 | LOB domain-containing protein 18 | ||||



