 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_4AL_5062917C4.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 941aa MW: 103212 Da PI: 5.8655 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128 | 3.7e-40 | 133 | 210 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC adls ak+yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Traes_4AL_5062917C4.1 133 ACQVEGCCADLSAAKDYHRRHKVCEMHAKANTAVVGNTVQRFCQQCSRFHLLQEFDEGKRSCRRRLAGHNKRRRKTRP 210
5**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.5E-33 | 126 | 195 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.291 | 131 | 208 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.75E-37 | 132 | 211 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.8E-29 | 134 | 207 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 4.35E-7 | 721 | 826 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.4E-6 | 723 | 828 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.75E-8 | 726 | 825 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 941 aa Download sequence Send to blast |
GELEQARREK RVFGWDLNDW SWDSERFVAT PVPAAVGNGL SLNSSPSSSE EAGAEVSRNG 60 NVRGDFDKRK RVVVIHDDDE KDEGPVGSSN NVLSLRIGGN SVAGGAVEDG GVNEEDRNGK 120 KIRVQGGSSS GPACQVEGCC ADLSAAKDYH RRHKVCEMHA KANTAVVGNT VQRFCQQCSR 180 FHLLQEFDEG KRSCRRRLAG HNKRRRKTRP ENAVGGTPIE EKVSSYLLLS LLGICANLNS 240 DNAEHLQGQE LLSNLWRNLG NVAKSLDPKE LCKLLETCQS MQNRSNAGTS EAANALVNTA 300 AAEAAGPSNS KAPFANGGEC GQTSSAVVPV QSNATMVAAT ETPACKFRNF DLNDTCNDME 360 GFEDGSNCPS WIRQDSTQSP PQTSGNSDST SAQSLSSSNG DAQCRTDKIV FKLFEKVPSE 420 LPPILRSQIL GWLSSSPTDI ESHIRPGCII LTVYLRLVES SWRELSENMS VYLDKLSSSS 480 ADNFWTSGLV FVMVRRQIAF MHNGQVMLDR PLAPNSHHYC KVLCVSPVAA PYSATVNFRV 540 EGFNLVSSSS RLICSIEGRC IFEEDTAIMA DDAEDEDIEY LNFCCSLPDT RGRGFIEVED 600 SGFSNGFFPF IVAEQNVCSE VCELESTFKS SSLEQPDKDN AMNQALEFLH ELGWLLHRVN 660 IISKHDKVEP PVPAFNLLRF RNLGIFAMER EWCAVTKMLL DLLFDGFVDA GLQSPKEVVL 720 SENLVHSAVR GKSARMVRFL LTYKPNKNLK ETAETYLFRP DARGPSAFTP LHIAASTSDA 780 EDVLDALTDD PGLVGLNAWR NARDEIGFTP EDYARQRGND AYINLVQKKI DRHLGKGHVV 840 LGVPSSMCPG ITDGLKAGDI SLEICKAMPM TTTSAARCNI CSRQAKMYPN SFARTFLYRP 900 AMFTVMGVAV ICVCVGILLH TLPKVYAAPN FRWELLERGA M |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-31 | 127 | 207 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
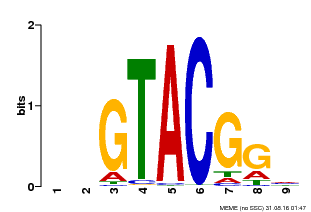
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF447880 | 0.0 | KF447880.1 Triticum aestivum squamosa promoter-binding-like protein 6 (SPL6) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020169994.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A446RFC7 | 0.0 | A0A446RFC7_TRITD; Uncharacterized protein | ||||
| STRING | Traes_4AL_5062917C4.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



