 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_2BL_569371098.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 433aa MW: 46574.9 Da PI: 9.6259 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 42 | 2.2e-13 | 94 | 154 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng...kr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++gr++A+++d ++ g k + + g ++ +e+Aa+a+++a++k++g
Traes_2BL_569371098.2 94 SQYRGVTRHRWTGRYEAHLWDnSCKkEGqtrKGrQVHFTGGYDMEEKAARAYDQAALKYWG 154
78*******************98888666863355555688******************98 PP
| |||||||
| 2 | AP2 | 52.4 | 1.4e-16 | 199 | 248 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GV+++++ grW A+I +s +k +lg+fgt eeAa+a++ a+ k++g
Traes_2BL_569371098.2 199 YRGVTRHHQHGRWQARIGRVSG---NKDLYLGTFGTQEEAAEAYDIAAIKFRG 248
9***************999754...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.01E-15 | 94 | 162 | No hit | No description |
| Pfam | PF00847 | 7.2E-10 | 94 | 154 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.25E-14 | 94 | 163 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 16.595 | 95 | 162 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 9.3E-14 | 95 | 163 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.7E-23 | 95 | 168 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.4E-6 | 96 | 107 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.16E-22 | 197 | 258 | No hit | No description |
| SuperFamily | SSF54171 | 9.81E-18 | 197 | 257 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 18.979 | 198 | 256 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.2E-18 | 198 | 256 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.1E-32 | 198 | 262 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 2.9E-11 | 199 | 248 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.4E-6 | 238 | 258 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 433 aa Download sequence Send to blast |
MANWVQVARG ATAYATAENV LSAAADRQQH LHHHPLALSM SSAGSLSSCV TAGAEYGGVV 60 ATVDGGRKRG GATAGQKQPV HHRKSIDTFG QRTSQYRGVT RHRWTGRYEA HLWDNSCKKE 120 GQTRKGRQVH FTGGYDMEEK AARAYDQAAL KYWGPSTHIN FPLEDYQQEL EEMKNMTRQE 180 YVAHLRRKSS GFSRGASMYR GVTRHHQHGR WQARIGRVSG NKDLYLGTFG TQEEAAEAYD 240 IAAIKFRGLN AVTNFDITRY DVDKIMASNT LLPGELARRN KDANAAPLPL PAPDDCAASA 300 LVPVSTPGTD TGGSGQHRHH DVMSSGEAFS ALHDLVTVDG HTAQGGNGAH VHMSMSGASS 360 LVTSLSNSRE GSPDRGGGLS MLFAKPPQQP ATTTAASPKL MSTLAPLGSW ASSARPAAVS 420 IAHMPMFAAW SDA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as positive regulator of adventitious (crown) root formation by promoting its initiation. Promotes adventitious root initiation through repression of cytokinin signaling by positively regulating the two-component response regulator RR1. Regulated by the auxin response factor and transcriptional activator ARF23/ARF1. {ECO:0000269|PubMed:21481033}. | |||||
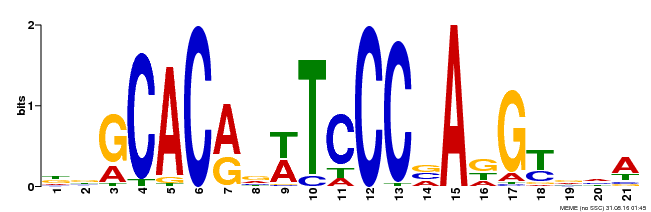
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by auxin. {ECO:0000269|PubMed:21481033}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT088263 | 1e-178 | BT088263.1 Zea mays full-length cDNA clone ZM_BFc0105N21 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020155443.1 | 0.0 | AP2-like ethylene-responsive transcription factor ANT isoform X2 | ||||
| Swissprot | Q84Z02 | 0.0 | CRL5_ORYSJ; AP2-like ethylene-responsive transcription factor CRL5 | ||||
| TrEMBL | A0A446M9E3 | 0.0 | A0A446M9E3_TRITD; Uncharacterized protein | ||||
| STRING | Traes_2BL_569371098.2 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP948 | 38 | 133 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37750.1 | 1e-127 | AP2 family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



