 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_1AL_6B108514B.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 192aa MW: 21807.5 Da PI: 10.5521 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 94 | 6.9e-30 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien + rqvtfskRrng+lKKA+ELSvLCdaeva+++fs++g+lye++s
Traes_1AL_6B108514B.1 9 KRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALVVFSPRGRLYEFAS 59
79***********************************************86 PP
| |||||||
| 2 | K-box | 70.9 | 4e-24 | 80 | 172 | 7 | 99 |
K-box 7 ksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaL 94
++ + + ++ + ++ L k++e L+ ++R++lGe+L+ +s +eL+ Le ++eksl+ iR kK++ll +qi +l++ke+ l ++n++L
Traes_1AL_6B108514B.1 80 NKIVQPDIQQVKADALSLAKKLEALEDTKRKILGENLGGCSTEELHFLEGKIEKSLRVIRGKKTQLLEQQIANLKEKERTLLKDNEDL 167
4477788999****************************************************************************** PP
K-box 95 rkkle 99
r k++
Traes_1AL_6B108514B.1 168 RGKVM 172
*9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 31.455 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.3E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 4.41E-39 | 3 | 76 | No hit | No description |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 7.2E-31 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 7.98E-31 | 3 | 83 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 6.1E-27 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 7.2E-31 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 7.2E-31 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 5.5E-25 | 86 | 171 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 13.161 | 87 | 177 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000060 | Biological Process | protein import into nucleus, translocation | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010077 | Biological Process | maintenance of inflorescence meristem identity | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008134 | Molecular Function | transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 192 aa Download sequence Send to blast |
MVRGKTQMKR IENATSRQVT FSKRRNGLLK KAFELSVLCD AEVALVVFSP RGRLYEFASA 60 TSLQKSIDRY KAYTKDTVNN KIVQPDIQQV KADALSLAKK LEALEDTKRK ILGENLGGCS 120 TEELHFLEGK IEKSLRVIRG KKTQLLEQQI ANLKEKERTL LKDNEDLRGK VMTLNKLLLP 180 PSHHIKAFLT LT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3mu6_A | 2e-18 | 3 | 73 | 2 | 71 | Myocyte-specific enhancer factor 2A |
| 3mu6_B | 2e-18 | 3 | 73 | 2 | 71 | Myocyte-specific enhancer factor 2A |
| 3mu6_C | 2e-18 | 3 | 73 | 2 | 71 | Myocyte-specific enhancer factor 2A |
| 3mu6_D | 2e-18 | 3 | 73 | 2 | 71 | Myocyte-specific enhancer factor 2A |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00076 | ChIP-chip | Transfer from AT2G45660 | Download |
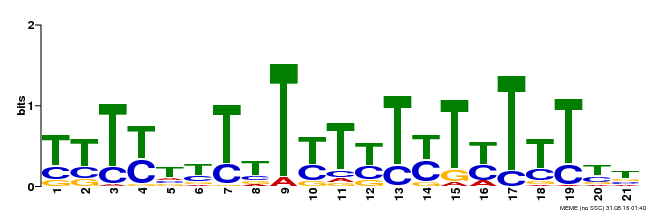
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM502862 | 0.0 | AM502862.1 Triticum aestivum mRNA for MIKC-type MADS-box transcription factor WM1B (WM1B gene). | |||
| GenBank | DQ512363 | 0.0 | DQ512363.1 Triticum aestivum MADS-box transcription factor TaAGL7 (AGL7) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020161859.1 | 1e-118 | MADS-box transcription factor 56-like isoform X2 | ||||
| Swissprot | A2Z9Q7 | 9e-95 | MAD56_ORYSI; MADS-box transcription factor 56 | ||||
| TrEMBL | A0A446IYC8 | 1e-136 | A0A446IYC8_TRITD; Uncharacterized protein | ||||
| STRING | Traes_1AL_6B108514B.1 | 1e-137 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1413 | 33 | 79 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45660.1 | 2e-63 | AGAMOUS-like 20 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



