 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRAES3BF096900180CFD_t1 | ||||||||
| Common Name | TRAES_3BF096900180CFD_c1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 300aa MW: 31509.6 Da PI: 10.8475 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 45.1 | 2.3e-14 | 116 | 160 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+ +++ + ++lG+g+W+ I+r++ ++Rt+ q+ s+ qky
TRAES3BF096900180CFD_t1 116 PWTEDEHRRFLGGLEKLGKGDWRGISRHFVTTRTPTQVASHAQKY 160
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50158 | 8.581 | 3 | 18 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 19.092 | 109 | 165 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.42E-17 | 110 | 165 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-18 | 112 | 164 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 9.9E-11 | 113 | 159 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.2E-10 | 113 | 163 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.4E-11 | 116 | 160 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.46E-9 | 116 | 161 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 300 aa Download sequence Send to blast |
MARKCSSCGH NGHNSRTCSG HRGMESGGGG LRLFGVQLQV GAAPLKKSFS MECLSSSASA 60 YNAAAAAVGV AASNSSSSVS SSSSLVSVEE SPEKMGQGYL SDGLMGRAQE RKKGVPWTED 120 EHRRFLGGLE KLGKGDWRGI SRHFVTTRTP TQVASHAQKY FLRQAGLAQK KRRSSLFDVV 180 EKNGDRGTTE RRHRLKPDAT SSVDAMGLTF PALSLGASRP RPDAALPPCL TLMPSCSPPS 240 SSASRAPKLP PSLGLVANAN PPRQAPDLEL KISSTAARKT DKQAGAAAGS SPFFGTIRVT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor (PubMed:23888064, PubMed:24806884). Direct regulator of the transcription of peroxidase (Prxs) and reactive oxygen species (ROS)-related genes via the recognition of 5'-ATCACA-3' motif (PubMed:24806884). Binds to 5'-TATCCA-3' motif (TA box) and represses the activity of corresponding promoters (e.g. sugar response genes) (PubMed:25920996). Regulates hypocotyl elongation in response to darkness by enhancing auxin accumulation in a phytochrome-interacting factor (PIF) proteins-dependent manner. Promotes lateral roots formation (PubMed:23888064). Promotes cell expansion during leaves development via the modulation of cell wall-located Prxs (PubMed:24806884). Plays a critical role in developmentally regulated and dark-induced onset of leaf senescence by repressing the transcription of several genes involved in chloroplast function and responses to light and auxin. Promotes responses to auxin, abscisic acid (ABA), and ethylene (PubMed:25920996). {ECO:0000269|PubMed:23888064, ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. | |||||
| UniProt | Transcription repressor that binds to 5'-TATCCA-3' elements in gene promoters. Contributes to the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. Transcription repressor involved in a cold stress response pathway that confers cold tolerance. Suppresses the DREB1-dependent signaling pathway under prolonged cold stress. DREB1 responds quickly and transiently while MYBS3 responds slowly to cold stress. They may act sequentially and complementarily for adaptation to short- and long-term cold stress (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00565 | DAP | Transfer from AT5G56840 | Download |
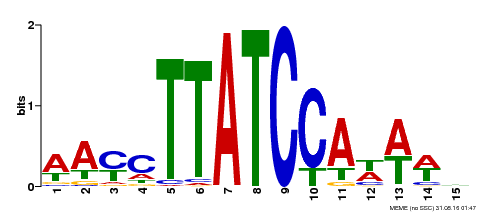
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by sucrose and gibberellic acid (GA) (PubMed:12172034). Induced by cold stress in roots and shoots. Induced by salt stress in shoots. Down-regulated by abscisic aci (ABA) in shoots (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. | |||||
| UniProt | INDUCTION: Slightly induced by CdCl(2) (PubMed:16463103). Accumulates in the dark (PubMed:23888064, PubMed:25920996). Diurnal expression pattern with maximal levels in the morning (at protein level). Specifically induced during leaf expansion (PubMed:24806884). Expressed in old and dark-treated leaves (PubMed:25920996). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:23888064, ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ624399 | 0.0 | KJ624399.1 Triticum aestivum MYBSM151 mRNA, complete cds. | |||
| GenBank | KP330479 | 0.0 | KP330479.1 Triticum aestivum MYB transcription factor SM151 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020153631.1 | 1e-171 | uncharacterized protein LOC109738954 | ||||
| Swissprot | Q7XC57 | 5e-49 | MYBS3_ORYSJ; Transcription factor MYBS3 | ||||
| Swissprot | Q9LVS0 | 1e-48 | KUA1_ARATH; Transcription factor KUA1 | ||||
| TrEMBL | A0A077RX78 | 0.0 | A0A077RX78_WHEAT; Uncharacterized protein | ||||
| STRING | Traes_3B_B45238368.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4231 | 38 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56840.1 | 4e-44 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



