 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRAES3BF001100080CFD_t1 | ||||||||
| Common Name | TRAES_3BF001100080CFD_c1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 796aa MW: 89098.7 Da PI: 6.483 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.2 | 6.6e-23 | 126 | 227 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-E CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDf 79
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d++g W++++i+r++++r++l++GW+ Fv++++L +gD
TRAES3BF001100080CFD_t1 126 FCKTLTASDTSTHGGFSVLRRHADEClppldmtQSPPT--QELVAKDLHGMDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDA 209
99*****************************7555544..58******************************************** PP
EEEEE-SSSEE..EEEEE-S CS
B3 80 vvFkldgrsefelvvkvfrk 99
++F + +++el+v+v+r+
TRAES3BF001100080CFD_t1 210 FIFL--RGESGELRVGVRRA 227
****..449999*****996 PP
| |||||||
| 2 | Auxin_resp | 111.2 | 9.9e-37 | 252 | 333 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF+++++++++++k+++s+GmRf+m+fe+e+++e+r++Gt+vg ++ld+ Wp+S+WrsLk
TRAES3BF001100080CFD_t1 252 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSIGMRFRMRFEGEEAPEQRFTGTIVGSENLDQ-LWPESNWRSLK 333
89******************************************************************999.9********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.27E-39 | 119 | 255 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.4E-39 | 119 | 241 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.04E-19 | 124 | 226 | No hit | No description |
| PROSITE profile | PS50863 | 11.702 | 126 | 228 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 9.5E-22 | 126 | 228 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 9.5E-21 | 126 | 227 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.2E-33 | 252 | 333 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.0E-12 | 659 | 759 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 1.57E-12 | 669 | 752 | No hit | No description |
| PROSITE profile | PS51745 | 24.728 | 675 | 768 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 796 aa Download sequence Send to blast |
MPPAAMAPPQ APSAGDPLYD ELWHACAGPL VTVPRVGDLV FYFPQGHIEQ VEASMNQVAG 60 NQMRLYDLPS KLLCRVINVE LKAEADTDEV YAQVMLMPEP EQNEMAVDKS TSTTGATPPR 120 PAVRSFCKTL TASDTSTHGG FSVLRRHADE CLPPLDMTQS PPTQELVAKD LHGMDWRFRH 180 IFRGQPRRHL LQSGWSVFVS SKRLVAGDAF IFLRGESGEL RVGVRRAMRQ LPNVPSSVIS 240 SHSMHLGVLA TAWHAINTKS MFTVYYKPRT SPSEFIIPYD QYMESVKNNY SIGMRFRMRF 300 EGEEAPEQRF TGTIVGSENL DQLWPESNWR SLKVRWDEPS TIPRPDRVSP WKIEPASSPP 360 VNPLPLSRVK RPRPNVPPVS PESSVLTKEG ATKIDMDSAQ AQQRNQNNMV LQGQEHMTLR 420 TNNLTASNDS DATVQKPMMW SPSPNIGKNH ASAFQQRPSM DNWMQLGRCD ASSGAQSFGD 480 SQGFFMQTFD EAPNRHGSFK NQFQDHSSAR HFSDPYTKMQ TEANEFHFWN SQSTVYGNPR 540 DQSQGFRFEE HPSNWLRQQQ FSPVEQPRVI RPHASIAPVD LEKAREGSGF KIFGFKVDTT 600 STPSNHLSSS MAAIHEPVLQ IQASASLTQL QHTHADCIPE LSVSTAGTTE NEKSIQQAPH 660 SSKDVQSKSH GASTRSCTKV HKQGVALGRS VDLSKFGDYD ELTAELDRMF EFDGELMSSN 720 KDWQIVYTDP EGDMMLVGDD PWEEFCNIVR KIFIYTKEEV QKMNSKSSAP RKEEGSGDAD 780 GANEKAHLAT SSHLDN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldx_A | 1e-167 | 16 | 358 | 17 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-167 | 16 | 358 | 17 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 369 | 373 | KRPRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
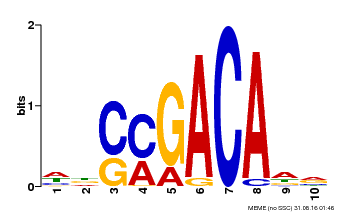
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK364852 | 0.0 | AK364852.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2028N02. | |||
| GenBank | AK368785 | 0.0 | AK368785.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2079M07. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020198715.1 | 0.0 | auxin response factor 4-like | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A077S779 | 0.0 | A0A077S779_WHEAT; Auxin response factor | ||||
| STRING | Traes_3B_615705174.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



