 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.9G526200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 322aa MW: 33564.6 Da PI: 6.1741 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.9 | 6.8e-20 | 29 | 78 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GVr++ +g+WvAeIr+p +kr+r +lg+f taeeAa a+++a+++l+g
Sevir.9G526200.1.p 29 EYRGVRQRT-WGKWVAEIREP---NKRTRLWLGSFATAEEAALAYDEAARRLYG 78
69****999.**********9...337*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.81E-28 | 28 | 88 | No hit | No description |
| Pfam | PF00847 | 7.1E-14 | 29 | 78 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.55 | 29 | 86 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.25E-21 | 29 | 87 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.0E-30 | 29 | 87 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.5E-36 | 29 | 92 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.2E-10 | 30 | 41 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.2E-10 | 52 | 68 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 322 aa Download sequence Send to blast |
MESYGGGRKR AWKKGPTRGK GGPQNAACEY RGVRQRTWGK WVAEIREPNK RTRLWLGSFA 60 TAEEAALAYD EAARRLYGPD AFLNLPHLRA SVTAAAAAAH QRLRWLPASA RGGAAVPAYG 120 LLNLNAQHNV HVIHQRLQEL KNGGAPPAKP PCGGGARAPH QLVVPPLPAS TTSPCSTVTN 180 GGGVAAHAAL PPPMSCFQAL EEAVATAAMT ADDDTEPCEG ACAGTDKPQL DLREFLQQIG 240 VLKTDEDGAT AKEGLHGDGD VAGAFGFGGN GEFDWDALAA DLNDIAGGHG GAVGVNGGFQ 300 MDDLHEVDQF GSCLPIPVWD V* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-16 | 30 | 86 | 6 | 63 | ATERF1 |
| 3gcc_A | 2e-16 | 30 | 86 | 6 | 63 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that binds to the DNA sequence 5'-[AG]CCGAC-3' of the cis-acting dehydration-responsive element (DRE). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00411 | DAP | Transfer from AT3G57600 | Download |
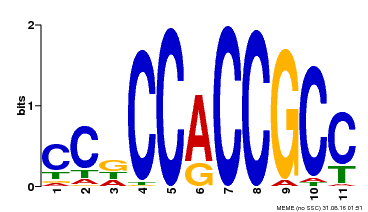
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.9G526200.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not induced by high-salt and drought stresses. {ECO:0000269|PubMed:20049613}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ728407 | 1e-166 | KJ728407.1 Zea mays clone pUT6712 AP2-EREBP transcription factor (EREB124) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004985522.1 | 0.0 | dehydration-responsive element-binding protein 2E | ||||
| Swissprot | Q10R18 | 1e-134 | DRE2E_ORYSJ; Dehydration-responsive element-binding protein 2E | ||||
| TrEMBL | A0A368SX52 | 0.0 | A0A368SX52_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.Ia04341.1.p | 1e-168 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9575 | 29 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G57600.1 | 4e-47 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.9G526200.1.p |



