 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.5G410700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 298aa MW: 32707.2 Da PI: 8.4344 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 162.9 | 1.2e-50 | 10 | 134 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdke 91
lppGfrFhPtdeelv++yL+++++g ++ + +i+e+d+yk++Pw+Lp+++ +ekewyfFs+rd+ky++g+r+nra+ +gyWkatg dk+
Sevir.5G410700.1.p 10 LPPGFRFHPTDEELVTHYLCRRCAGLPIAV-PIIAEIDLYKFDPWQLPRMALYGEKEWYFFSPRDRKYPNGSRPNRAAGTGYWKATGADKP 99
79**************************99.88***************7666789************************************ PP
NAM 92 vlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
v + + +kk Lvfy g+apkgekt+W+mheyrl
Sevir.5G410700.1.p 100 VGT--PKPLAIKKALVFYAGKAPKGEKTNWIMHEYRL 134
*99..7789**************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 7.72E-61 | 8 | 160 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 59.351 | 10 | 160 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.5E-26 | 11 | 134 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 298 aa Download sequence Send to blast |
MSGGGQDLQL PPGFRFHPTD EELVTHYLCR RCAGLPIAVP IIAEIDLYKF DPWQLPRMAL 60 YGEKEWYFFS PRDRKYPNGS RPNRAAGTGY WKATGADKPV GTPKPLAIKK ALVFYAGKAP 120 KGEKTNWIMH EYRLADVDRS ARKKNSLRLD DWVLCRIYNK KGGLEKPPVA AGDRKPALLA 180 PASAGSPPEQ KPFVAAPGGL PPAFADLAAY YDRPSDSMPR LHADSSCSEQ VLSPEQQFAG 240 CDREVQSQPK ISEWERTFAS DPVNPAGSLL DPVGHGGDGI GGDPLLQDIL MYWGKPF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 2e-78 | 7 | 166 | 12 | 174 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the promoter of the stress response gene LEA19. Involved in tolerance to abiotic stresses (PubMed:20632034). Transcription activator involved in response to abiotic and biotic stresses. Involved in drought and salt stress responses, and defense response to the rice blast fungus (PubMed:17587305). Transcription activator involved tolerance to cold and salt stresses (PubMed:18273684). Transcription activator involved in tolerance to drought stress. Targets directly and activates genes involved in membrane modification, nicotianamine (NA) biosynthesis, glutathione relocation, accumulation of phosphoadenosine phosphosulfate and glycosylation in roots (PubMed:27892643). Controls root growth at early vegetative stage through chromatin modification and histone lysine deacytaltion by HDAC1 (PubMed:19453457). {ECO:0000269|PubMed:17587305, ECO:0000269|PubMed:18273684, ECO:0000269|PubMed:19453457, ECO:0000269|PubMed:20632034, ECO:0000269|PubMed:27892643}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
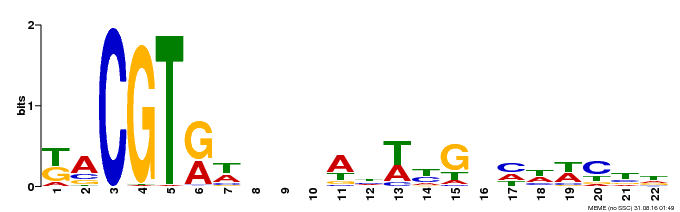
| Motif ID | Method | Source | Motif file |
| MP00121 | DAP | Transfer from AT1G01720 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.5G410700.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by drought stress, salt stress, cold stress and abscisic acid (ABA) (PubMed:20632034, PubMed:27892643). Induced by methyl jasmonate (PubMed:20632034, PubMed:11332734). Induced by infection with the rice blast fungus Magnaporthe oryzae (PubMed:11332734). {ECO:0000269|PubMed:11332734, ECO:0000269|PubMed:20632034, ECO:0000269|PubMed:27892643}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY742218 | 0.0 | AY742218.1 Saccharum officinarum NAC23 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004970815.1 | 0.0 | NAC domain-containing protein 48 | ||||
| Swissprot | Q7F2L3 | 1e-170 | NAC48_ORYSJ; NAC domain-containing protein 48 | ||||
| TrEMBL | A0A368RE84 | 0.0 | A0A368RE84_SETIT; Uncharacterized protein | ||||
| STRING | Si002413m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3417 | 37 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01720.1 | 1e-118 | NAC family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.5G410700.1.p |



