| Signature Domain? help Back to Top |
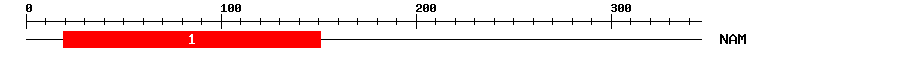 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | NAM | 169.3 | 1.2e-52 | 19 | 151 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdke 91
lppGfrF P+dee++++yLk+kv+++++++ ++i evd++++ePw+L k+k ++kewyfF +d+ky+tgkr+nratk gyWkatgkd+e
Sevir.5G071600.1.p 19 LPPGFRFYPSDEEIITFYLKPKVHQSNFSC-TAIGEVDLNRIEPWELSGKAKMGDKEWYFFYLKDRKYPTGKRTNRATKGGYWKATGKDRE 108
79****************************.88**************999999************************************** PP
NAM 92 vlskkgel.....vglkktLvfykgrapkgektdWvmheyrle 129
++++ +++ vg+kktLvfykgrap+g+ktdWvmhe+rle
Sevir.5G071600.1.p 109 IYRAAKKDelpllVGMKKTLVFYKGRAPTGKKTDWVMHEFRLE 151
***8555578999***************************985 PP
|
| Functional Description ? help
Back to Top |
| Source |
Description |
| UniProt | Binds to the promoter regions of genes involved in chlorophyll catabolic processes, such as NYC1, SGR1, SGR2 and PAO. {ECO:0000269|PubMed:27021284}. |
| UniProt | Transcription activator that binds to DNA in promoters of target genes on a specific bipartite motif 5'-[ACG][CA]GT[AG](5-6n)[CT]AC[AG]-3' (PubMed:23340744). Promotes lateral root development (PubMed:16359384). Triggers the expression of senescence-associated genes during age-, salt- and dark-induced senescence through a regulatory network that may involve cross-talk with salt- and H(2)O(2)-dependent signaling pathways (PubMed:9351240, PubMed:15295076, PubMed:20113437, PubMed:21303842). Regulates also genes during seed germination (PubMed:20113437). Regulates positively aging-induced cell death (PubMed:19229035). Involved in age-related resistance (ARR) against Pseudomonas syringae pv. tomato and Hyaloperonospora arabidopsidis (PubMed:19694953). Antagonizes GLK1 and GLK2 transcriptional activity, shifting the balance from chloroplast maintenance towards deterioration during leaf senescence (PubMed:23459204). Promotes the expression of senescence-associated genes, including ENDO1/BFN1, SWEET15/SAG29 and SINA1/At3g13672, during senescence onset (PubMed:23340744). {ECO:0000269|PubMed:15295076, ECO:0000269|PubMed:16359384, ECO:0000269|PubMed:19229035, ECO:0000269|PubMed:19694953, ECO:0000269|PubMed:20113437, ECO:0000269|PubMed:21303842, ECO:0000269|PubMed:23340744, ECO:0000269|PubMed:23459204, ECO:0000269|PubMed:9351240}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: High levels during senescence (e.g. age-, salt- and dark-related) (PubMed:19229035, PubMed:20113437, PubMed:21511905, PubMed:22930749). By salt stress in an ethylene- and auxin-dependent manner (PubMed:16359384, PubMed:19608714, PubMed:20113437, PubMed:20404534). Induced by H(2)O(2) (PubMed:20404534). Accumulates in response to abscisic acid (ABA), ethylene (ACC) and auxin (NAA) (PubMed:16359384, PubMed:19608714). Repressed by high auxin (IAA) levels (PubMed:21511905). Age-related resistance (ARR)-associated accumulation (PubMed:19694953). Repressed by miR164 (PubMed:19229035). {ECO:0000269|PubMed:16359384, ECO:0000269|PubMed:19229035, ECO:0000269|PubMed:19608714, ECO:0000269|PubMed:19694953, ECO:0000269|PubMed:20113437, ECO:0000269|PubMed:20404534, ECO:0000269|PubMed:21511905, ECO:0000269|PubMed:22930749}. |
| UniProt | INDUCTION: Repressed by the microRNA miR164. {ECO:0000269|PubMed:15294871, ECO:0000269|PubMed:17098808}. |




