 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400027896 | ||||||||
| Common Name | LOC102604886 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 518aa MW: 57437.2 Da PI: 7.5302 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 54 | 3.1e-17 | 335 | 381 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+ellP++ +K +Ka++L +A+eY+ksLq
PGSC0003DMP400027896 335 VHNLSERRRRDRINEKMKALQELLPHS-----TKTDKASMLDEAIEYLKSLQ 381
6*************************9.....9******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 18.476 | 331 | 380 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.71E-18 | 334 | 385 | No hit | No description |
| SuperFamily | SSF47459 | 3.4E-20 | 334 | 408 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.6E-20 | 335 | 389 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.3E-14 | 335 | 381 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.0E-18 | 337 | 386 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 518 aa Download sequence Send to blast |
MNPCLPEWNI ETELPVPHQK KPMGFDHELV ELLWRNGEVV LHNQTHKKQP GYDPNECRQF 60 NKYDQPTIRV AGNQTNLIQD DETVSWLNCP IYDSFDKEFC SPFLSDISTN PHLGEEPDKS 120 IRQSEDNNKV FKLDPLEINH VLPQSHHSGF NPNPMPPPRF HNFGSAQQKH HIVGGNQKGV 180 NFPPPIRSSN VQLGGKEARS NLMLQDIKEG SVMTVGSSHC GSNQVDTSRF SSSANRGLSA 240 AMITDYTGKI SPQSDTMDRD TFEPANTSSS SGRSGSSYAR ACNQSTATNS QGHKRKSRDG 300 EEPECQSKAD ELESAGGNKA AQKSGTARRS RAAEVHNLSE RRRRDRINEK MKALQELLPH 360 STKTDKASML DEAIEYLKSL QMQLQMMWMG SGMASMMFPG VQHYMSRMGM GMGPPSVPSM 420 HNAMHLARLP LVDPAIPLTQ AAPNNQAAAM CQNSMLNQVN YQRHLQNPNF PEQYASYMGF 480 HPLQGTSQPI NIFGLGSHTA QRSQQLPHPT NSNAPAT* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 339 | 344 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
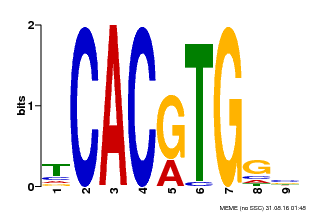
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400027896 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013582 | 0.0 | BT013582.1 Lycopersicon esculentum clone 132330F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006357888.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_015169146.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_015169147.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| TrEMBL | M1BAZ9 | 0.0 | M1BAZ9_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400041156 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 1e-44 | phytochrome interacting factor 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400027896 |
| Entrez Gene | 102604886 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



