 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400017819 | ||||||||
| Common Name | LOC102591830 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 366aa MW: 40693 Da PI: 8.8901 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.9 | 2e-32 | 231 | 286 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++qLGG+++AtPk+i+++m+v+gLt+++vkSHLQkYRl+
PGSC0003DMP400017819 231 KQRRCWSPELHRRFVDALHQLGGAQVATPKQIRDIMQVDGLTNDEVKSHLQKYRLH 286
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 9.1E-30 | 228 | 289 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.929 | 228 | 288 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.97E-17 | 229 | 289 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.7E-26 | 231 | 286 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.5E-7 | 233 | 284 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010092 | Biological Process | specification of organ identity | ||||
| GO:0010629 | Biological Process | negative regulation of gene expression | ||||
| GO:0048449 | Biological Process | floral organ formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 366 aa Download sequence Send to blast |
MGSNSKEMNI DLNFVYVPKL ISDVLTEVSA MDDISKKLTK LNQFLVPLEE ELRKTEAFKR 60 ELPMCMLLLK DAIERLKAEA LLYKEKDKSP VMEEFIPLKK GNSDESGRVK KSNDLSDKKN 120 WMSSAQLWST PVQYESLKSS GVEEKAREKQ YQLGACKLKN ERGAFLPFQG QALKEGKNGL 180 AVKDLSLSMA VTVGEGEKGQ KPIDVSVKRE NGPSNSNGCG VFSQDKSQHR KQRRCWSPEL 240 HRRFVDALHQ LGGAQVATPK QIRDIMQVDG LTNDEVKSHL QKYRLHVRRV PASGCSWSNM 300 DEIGESSKTN GTQSGSPEGP LHFTGSGSAK GVSINEEEDN KSESYNWNGQ LQKSIEGSTI 360 RSSSH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-13 | 231 | 284 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-13 | 231 | 284 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-13 | 231 | 284 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-13 | 231 | 284 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-13 | 231 | 284 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
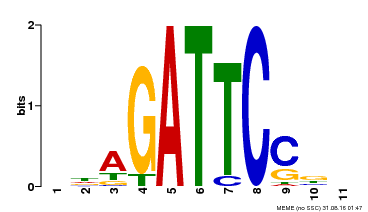
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400017819 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975441 | 1e-144 | HG975441.1 Solanum pennellii chromosome ch02, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006338223.1 | 0.0 | PREDICTED: uncharacterized protein LOC102591830 | ||||
| TrEMBL | M1AMN5 | 0.0 | M1AMN5_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400026096 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8497 | 22 | 29 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37180.1 | 2e-59 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400017819 |
| Entrez Gene | 102591830 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



