 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400016413 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 144aa MW: 16143.4 Da PI: 10.2802 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 106.1 | 1.9e-33 | 16 | 70 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWt +LH+rFv+av++LGG++kAtPk++l+lm++kgLtl+h+kSHLQkYRl
PGSC0003DMP400016413 16 KPRLRWTADLHDRFVDAVTKLGGPDKATPKSVLRLMGLKGLTLYHLKSHLQKYRL 70
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-32 | 13 | 71 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.455 | 13 | 73 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.53E-17 | 16 | 73 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.1E-25 | 16 | 71 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.4E-9 | 18 | 69 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 8.8E-10 | 116 | 138 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 144 aa Download sequence Send to blast |
MGYGYENGVV MTRDPKPRLR WTADLHDRFV DAVTKLGGPD KATPKSVLRL MGLKGLTLYH 60 LKSHLQKYRL GQQTKKQNTA DQNRENIGES FGQFSLHSSG PSITSSSMDG MQGEAPISEA 120 LRRQIDVQKR LHEQLEVHIK PFL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 9e-21 | 16 | 72 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 9e-21 | 16 | 72 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 9e-21 | 16 | 72 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 9e-21 | 16 | 72 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-20 | 16 | 72 | 2 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
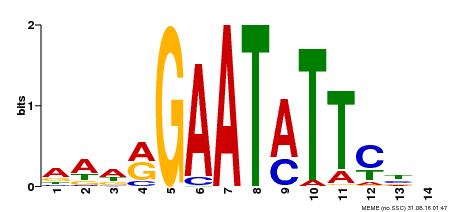
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400016413 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP009262 | 2e-60 | AP009262.1 Solanum lycopersicum genomic DNA, chromosome 8, clone: C08HBa0006A17, complete sequence. | |||
| GenBank | HG975447 | 2e-60 | HG975447.1 Solanum pennellii chromosome ch08, complete genome. | |||
| GenBank | HG975520 | 2e-60 | HG975520.1 Solanum lycopersicum chromosome ch08, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006352940.1 | 3e-98 | PREDICTED: myb family transcription factor APL isoform X2 | ||||
| Swissprot | C0SVS4 | 1e-59 | PHLB_ARATH; Myb family transcription factor PHL11 | ||||
| TrEMBL | M1AJD5 | 1e-102 | M1AJD5_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400024050 | 1e-87 | (Solanum tuberosum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45580.1 | 1e-58 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400016413 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



