 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.1001s0050.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 391aa MW: 42885 Da PI: 7.8911 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.3 | 1.3e-31 | 226 | 281 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
kpr++W+peLH+rF++a++qLGGs+ tPk+i+elmkv+gLt+++vkSHLQkYRl+
SapurV1A.1001s0050.1.p 226 KPRRCWSPELHRRFLHALQQLGGSHASTPKQIRELMKVDGLTNDEVKSHLQKYRLH 281
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 7.5E-29 | 223 | 284 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.976 | 223 | 283 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.3E-18 | 224 | 284 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.9E-26 | 226 | 281 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.6E-7 | 228 | 279 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 391 aa Download sequence Send to blast |
MDFAEKMQRC HEYVEALEAE RRKIQVFERE LPLCLELVTQ AIEACKRELS GTTADHNMNG 60 QPECSEQTSS EGPVLEEFIP IKRTHSSDDD ENDNNHDDDD DFQEQQSQNN NNNKRNKSNS 120 SNSNSDHKKK SDWLRSVQLW NQSPDPPPKQ DLPRKAAVTE VTRNGAGGAF QPFHREKSVG 180 KSSNQAIKAP TSVPASAISS TAGAVTGGTG GGGNKREDKE EGNQRKPRRC WSPELHRRFL 240 HALQQLGGSH ASTPKQIREL MKVDGLTNDE VKSHLQKYRL HTRRPNPTIH NNSNQQAPQF 300 VVVGGIWVPP AEYAAVAATI SGETSTISAA NGMCARKAAA PPAAPQNRQN TQSEHLRSEG 360 RGSRCERGSH SSNSPSTSSS THTTTTSPVF * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 7e-13 | 226 | 279 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 7e-13 | 226 | 279 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 7e-13 | 226 | 279 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 7e-13 | 226 | 279 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
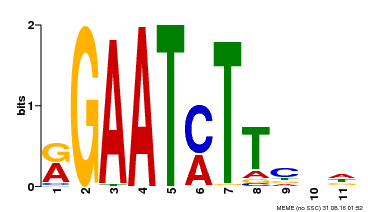
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.1001s0050.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024462049.1 | 0.0 | transcription factor HHO2 | ||||
| Swissprot | Q9FPE8 | 1e-108 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A2I4KKV8 | 0.0 | A0A2I4KKV8_POPTR; G2-like family protein | ||||
| STRING | POPTR_0010s13900.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3918 | 33 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 8e-93 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.1001s0050.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



