 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0767s0100.6.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 567aa MW: 65853.2 Da PI: 8.6409 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 21.6 | 7.1e-07 | 1 | 26 | 66 | 91 |
FAR1 66 lkvkkekdgkwevtkleleHnHelap 91
++vk++ dgkw+++++++eHnHel p
SapurV1A.0767s0100.6.p 1 MHVKRRPDGKWVIHSFVKEHNHELLP 26
89*********************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF10551 | 1.1E-31 | 124 | 215 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.322 | 404 | 440 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.4E-7 | 415 | 442 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF04434 | 7.6E-5 | 416 | 439 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 567 aa Download sequence Send to blast |
MHVKRRPDGK WVIHSFVKEH NHELLPAQAV SEQTRKMYAA MAQQFAEYKN VVGLKNDPKN 60 PFDKGWNLGL EAGETKILLD FFTRMQNMNS NFFYAVDLGE DQRLKNLIWA DAKSRHDYSN 120 FSDVVNFDTT YVRNKYKMPL ALFVGVNQHY QFMLLGCALV SDESAATYSW LMQTWLRAMG 180 GQTPKVIITD QDKVLKQVIS EVFPNAHHCF CLWNVLGKVS ENLGNVIKQN RNFMAKFDKC 240 IFRSWTENEF GKRWLKILDR FELRENEWMQ SLYEDREQWV PIYMRGAFLA GMSTALRSES 300 INSYFDKYVH KKTTVQEFVR QYESILQDRY EEEAKADSDT WNKQPALKSP SPLEKSVSGM 360 YTHAVFKKFQ VEVLGVVACH PKMESQDETS VSFRVQDLEK EQDFTVLWNQ MRLEVSCICR 420 LYEYKGYLCR HALVVLQMCQ QSAIPSQYIL KRWTKDAKSR HLVGEECEQV QSRVQRYNDL 480 CQRALKLSEE ASLSQESYSM AFRALEEAFG NCIGMNNSNK NLVEAGTSAT HGLLCIEDDN 540 QNRSVTKTIK KKNQTKKRKV GGQKQI* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
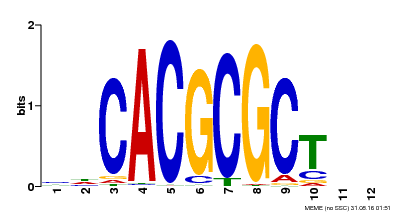
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0767s0100.6.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF148105 | 5e-74 | EF148105.1 Populus trichocarpa clone WS0127_J20 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024459456.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024459457.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024459458.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024459459.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024459460.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X1 | ||||
| Refseq | XP_024459461.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2K1ZVK4 | 0.0 | A0A2K1ZVK4_POPTR; Uncharacterized protein | ||||
| TrEMBL | B9HD60 | 0.0 | B9HD60_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0006s02150.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0767s0100.6.p |



