 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0739s0030.3.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 582aa MW: 63235.6 Da PI: 6.6856 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 90.3 | 1.7e-28 | 110 | 163 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
SapurV1A.0739s0030.3.p 110 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTREKVASHLQKYRL 163
79*******************.********************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50110 | 13.087 | 1 | 44 | IPR001789 | Signal transduction response regulator, receiver domain |
| SuperFamily | SSF52172 | 2.34E-6 | 5 | 47 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 1.5E-10 | 5 | 51 | No hit | No description |
| PROSITE profile | PS51294 | 10.737 | 107 | 166 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.27E-19 | 107 | 167 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 5.1E-30 | 109 | 168 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-24 | 110 | 163 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.9E-7 | 112 | 162 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 582 aa Download sequence Send to blast |
MSCAVMSADD GKNVVMKGVT HGACDYLIKP IRIEALKNIW QHVVRKKKKN EWKDLEQSGS 60 VEEGGDRQQK QSEDADYSSS ANEGSWKNSK RRKDEEEEVD ERDDTSTLKK PRVVWSVELH 120 QQFVAAVNQL GIDKAVPKKI LELMNVPGLT REKVASHLQK YRLYLRRLSG VSQHQSGMGN 180 SFISTQEATY GSLSSLNGLD LQTLAAASQI PAQSLATLQA AGLGRSTAKP RMPMPIVDQR 240 NLFSFENPKL RFGEGQQQHL NNGKQINLLH GIPTTMEPKQ LANMHHSAQS LGSMNMQLNA 300 HGDQSGSLLM QMSQQQSRGQ ILNEATSSQV PRLPSSIGQP IVNNALASGV LARNGLAESG 360 RGTGFNPVSQ SSSTMNFPLN TTAELTATSF ALGSAPGVPS LTSKGMFPEE ISSEIKGPGG 420 FIMPSYDIFS DLQHHRSHDW ELQNAGMTFN GSQQSNSLQS NVDVASSVLA HQGFSSSQSN 480 GQGRNISAVG KPMFSAGDAT DHVNAQSLGQ PLNNFFAENL VRVKTERVPD ASLQTTLFNE 540 QFGQEDLMSA LLKQQQQQQA GIGPAENEFD FDGYSLDNIP V* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-21 | 109 | 169 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
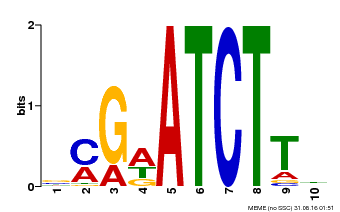
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0739s0030.3.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FN908139 | 0.0 | FN908139.1 Populus x canadensis mRNA for putative B-type response regulator 13 (rr13 gene), cultivar Dorskamp. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024466703.1 | 0.0 | two-component response regulator ARR2 isoform X2 | ||||
| Swissprot | Q9ZWJ9 | 1e-157 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A2K1YLL3 | 0.0 | A0A2K1YLL3_POPTR; Two-component response regulator | ||||
| STRING | POPTR_0008s22990.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-145 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0739s0030.3.p |



