 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0586s0110.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 826aa MW: 89446.5 Da PI: 6.0837 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.4 | 8e-21 | 118 | 173 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l L++rqVk+WFqNrR+++k
SapurV1A.0586s0110.1.p 118 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLCLETRQVKFWFQNRRTQMK 173
688999***********************************************999 PP
| |||||||
| 2 | START | 198.1 | 3.9e-62 | 328 | 552 | 1 | 205 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S. CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla. 77
ela++a++elvk+a+ +ep+W +s+ e+ n +e+ ++f++ + + +ea+r++g+v+ ++ lve+l+d++ +W e+++
SapurV1A.0586s0110.1.p 328 ELALAAMDELVKMAQTDEPLWIRSYeggrEIMNHEEYSRTFTPCIGmkpsgFVSEASRETGMVIINSLALVETLMDSN-RWAEMFPc 413
5899************************99***********9988899******************************.******** PP
...EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEE CS
START 78 ...kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgil 153
+ +t++vi +g g lqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + +s+v +++lpSg++
SapurV1A.0586s0110.1.p 414 viaRTSTTDVIANGmggtrnGSLQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGaPPSFVNCRRLPSGCV 500
*****************************************************************99999999************** PP
EEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 154 iepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+++++ng+skvtw+eh+++++++ h+l+r+l++sg+ +ga++w atlqrq e
SapurV1A.0586s0110.1.p 501 VQDMPNGYSKVTWIEHAEYDESQTHQLYRPLISSGMGFGAQRWIATLQRQSE 552
*************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.07E-20 | 101 | 175 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-21 | 105 | 175 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 115 | 175 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.7E-17 | 116 | 179 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.05E-18 | 117 | 175 | No hit | No description |
| Pfam | PF00046 | 2.2E-18 | 118 | 173 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 150 | 173 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.838 | 319 | 556 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.33E-34 | 321 | 553 | No hit | No description |
| CDD | cd08875 | 6.40E-123 | 323 | 552 | No hit | No description |
| SMART | SM00234 | 2.3E-46 | 328 | 553 | IPR002913 | START domain |
| Pfam | PF01852 | 8.0E-54 | 328 | 552 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.75E-23 | 581 | 751 | No hit | No description |
| SuperFamily | SSF55961 | 2.75E-23 | 778 | 818 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 826 aa Download sequence Send to blast |
MSFGGFLDNT SPGGGGARIV ADIPFNNNSM PTGGAIIAHP RLLSPSITKP MFNSPGLSLA 60 LQQPNIDGQG DITRMSENFE TSVGRRSREE EHESRSGSDN MDGASGDDQD AADNPPRKKR 120 YHRHTPQQIQ ELEALFKECP HPDEKQRLEL SRRLCLETRQ VKFWFQNRRT QMKTQLERHE 180 NSLLRQENDK LRAENMSIRD TMRNPMCSNC GGPAIIGDIS LEEQHLRIEN ARLKDELDRV 240 CALAGKFLGR PISSLASSLG PPMPNSSLEL GVGSNGFAGL STVAATLPLG PDFVGGISGA 300 LPLLTQARPA TTGVTGIGRS LERSMFLELA LAAMDELVKM AQTDEPLWIR SYEGGREIMN 360 HEEYSRTFTP CIGMKPSGFV SEASRETGMV IINSLALVET LMDSNRWAEM FPCVIARTST 420 TDVIANGMGG TRNGSLQLMH AELQVLSPLV PVREVNFLRF CKQHAEGVWA VVDVSIDTIR 480 ETSGAPPSFV NCRRLPSGCV VQDMPNGYSK VTWIEHAEYD ESQTHQLYRP LISSGMGFGA 540 QRWIATLQRQ SECLAILMSS NVPSRDHTAI TASGRRSMLK LAQRMTDNFC AGVCASTVHK 600 WNKLNAGNVD EDVRVMTRKS VDDPGEPPGI VLSAATSVWL PVSPQRLFDF LRDERLRSEW 660 DILSNGGPMQ EMAHIAKGQD HGNCVSLLRA SAMNANQSSM LILQETCIDA AGSLVVYAPV 720 DIPAMHVVMN GGDSAYVALL PSGFAIVPDG PGSLGAPAAN GGPTANNNGN GGGPERVSGS 780 LLTVAFQILV NSLPTAKLTV ESVETVNNLI SCTVQKIKAA LQCES* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
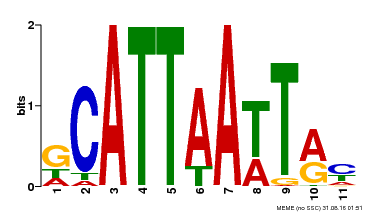
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0586s0110.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002320755.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | B9IC55 | 0.0 | B9IC55_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0014s07130.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0586s0110.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



