 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0118s0260.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 430aa MW: 47501.5 Da PI: 6.546 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 87.9 | 9.4e-28 | 155 | 208 | 2 | 56 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
++++WtpeLH+rFv+aveqL G +kA+P++ilelm++++Lt+++++SHLQkYR++
SapurV1A.0118s0260.2.p 155 KKVDWTPELHRRFVQAVEQL-GVDKAVPSRILELMGIDCLTRHNIASHLQKYRSH 208
689*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.806 | 151 | 210 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.6E-27 | 152 | 211 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.15E-18 | 154 | 211 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.3E-26 | 155 | 208 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.6E-8 | 157 | 206 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 430 aa Download sequence Send to blast |
MLAVSPLRNT AAAKDESQEQ MESYSTIFNG EFPDFSDGNL LESIDFDDLF VSIDDEQVLP 60 NLEMDPEILA EFSVSLSGGE ESDVNTSVSN ENVEDNIHDR KDEEDRFSGS GLDSKILSER 120 DESVVVNPVL PDKYGEKGRK SSAHHAKNNS NQGRKKVDWT PELHRRFVQA VEQLGVDKAV 180 PSRILELMGI DCLTRHNIAS HLQKYRSHRK HLQAREAEAA NWSQRRQMYG AAAAGGGGKR 240 DICAWHAFTM GFPPITRPMH HHFRPLHVWG HPSMDQSHMW PKHLAHSPSP PRPLPPPPTW 300 HPPPPPPPPP DPSYWHHHPH VRVPNGITPG TPCFPQPLAT RFPAPPVPGI PPYAMYKVDP 360 GIGLQTAGHP GRSPLVDFQP SKESVDAVIG DVLSKPWLPL PLGLKAPATD SVLVELQKQG 420 VPKIPPSFA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-15 | 152 | 206 | 3 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that functions with GLK2 to promote chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport. Acts in a cell-autonomous manner to coordinate and maintain the photosynthetic apparatus within individual cells. May function in photosynthetic capacity optimization by integrating responses to variable environmental and endogenous cues (PubMed:11828027, PubMed:12220263, PubMed:17533111, PubMed:18643989, PubMed:19376934, PubMed:19383092, PubMed:19726569). Prevents premature senescence (PubMed:23459204). {ECO:0000269|PubMed:11828027, ECO:0000269|PubMed:12220263, ECO:0000269|PubMed:17533111, ECO:0000269|PubMed:18643989, ECO:0000269|PubMed:19376934, ECO:0000269|PubMed:19383092, ECO:0000269|PubMed:19726569, ECO:0000269|PubMed:23459204}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
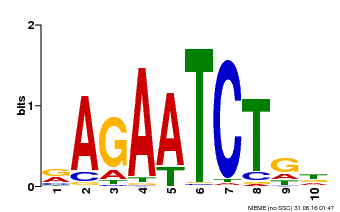
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0118s0260.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. Repressed by BZR2. {ECO:0000269|PubMed:12220263, ECO:0000269|PubMed:21214652}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC209002 | 1e-122 | AC209002.1 Populus trichocarpa clone POP040-E22, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024448608.1 | 0.0 | transcription activator GLK1-like | ||||
| Swissprot | Q9SIV3 | 1e-89 | GLK1_ARATH; Transcription activator GLK1 | ||||
| TrEMBL | B9HGR1 | 0.0 | B9HGR1_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0007s01230.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 4e-76 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0118s0260.2.p |



