 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0061s0540.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 371aa MW: 41162.7 Da PI: 8.7766 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.8 | 1.8e-31 | 73 | 126 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +Fv+ave+LGG+e+AtPk +l++m++k+L+++hvkSHLQ+YR+
SapurV1A.0061s0540.2.p 73 PRLRWTPDLHLCFVHAVERLGGQERATPKLVLQMMNIKDLSIAHVKSHLQMYRS 126
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-30 | 69 | 128 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.79E-14 | 71 | 128 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.4E-22 | 73 | 126 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.9E-9 | 74 | 125 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 371 aa Download sequence Send to blast |
MEGSDSFEGS KSSYSNKDVE QDEAGGEDDE VNNKISFKSR NEGESTSNSS VEENGKKATS 60 TGSVRQYVRS KMPRLRWTPD LHLCFVHAVE RLGGQERATP KLVLQMMNIK DLSIAHVKSH 120 LQMYRSMRND EPNQGQGLCF EGGDHQIYNL SQLPILQNFN QRSSCNLRFG DASWRGDHDR 180 QMYSPYTGGT GLNRFKHGLY GSVSERLVTG RNNHNSLNSG SSINIPSLNV QAATSRTHQF 240 LEGVRLFQVS RQNESRPGSM ESNFIAKLQE RSSGIDQKEC SNTTSSANKN WRTIQEMQKG 300 LKRKTLDSDC NLDLNLSLKL APKDDDRLQK CVVDGSLSPP LSSSSSSKLG TSMEGGRKHA 360 RMASTLDLTL * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-17 | 74 | 128 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-17 | 74 | 128 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-17 | 74 | 128 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-17 | 74 | 128 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
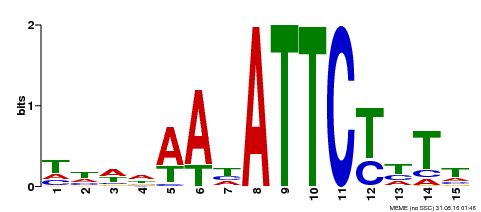
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0061s0540.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011035900.1 | 0.0 | PREDICTED: putative two-component response regulator ARR21 | ||||
| TrEMBL | B9HHT8 | 1e-176 | B9HHT8_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0010s19310.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 1e-54 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0061s0540.2.p |



