 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0014s0380.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 975aa MW: 108624 Da PI: 6.372 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.7 | 1.2e-41 | 149 | 226 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C +dls+ak+yhrrhkvCe+hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
SapurV1A.0014s0380.2.p 149 VCQVEDCGVDLSNAKDYHRRHKVCEMHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 226
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.5E-33 | 145 | 211 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.308 | 147 | 224 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 5.49E-39 | 148 | 229 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.5E-30 | 150 | 223 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 2.5E-6 | 742 | 857 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.76E-7 | 750 | 858 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 8.69E-8 | 751 | 858 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 975 aa Download sequence Send to blast |
METIFGGEAR HFYAMGSTDK REVGKRGLEW NLNDWKWDGD LFIASPLNPV PSTNVSRQLF 60 PLGVGTGVPA TGNSYSSSSF SDEVNLGVEK GKRELEKRRR VVVIDDDNLN DQETGGLNLK 120 LGGERDVGNW EGSSGKKTKL VGGGLSRAVC QVEDCGVDLS NAKDYHRRHK VCEMHSKASK 180 ALVGNVMQRF CQQCSRFHVL QEFDEGKRSC RRRLAGHNKR RRKTNPDTVG NGSSMNDDQT 240 SGYLLIRLLR ILSNMHSNRS NETTDQDLLT HLLRSLASHS VEHGGRNLFG LLQEPRDLST 300 SFGNSEQVMR MHDANGANIQ TTFSLKPSIP NNFATYSEVR ESTAVQVKMN NFDLNDIYVD 360 SDDGTEDIER SPAPVNARTS SLDCPSWVQK DSHQSSPPQT SRNSDSASAQ SPSSSGGEAQ 420 CRTDRIVFKL FGKEPNDFPL VLRAQILDWL SHSPTDIESY IRPGCIILTI YLHQAEAAWE 480 ELCCGLGSSL SRLLDVSEDT FWRTGWVYIR VQHQIAFVYN GQVVVDTSLP LTSNNCSKIL 540 SVKPIAITAT ERTEFLIKGI NLSRPATKLL CAVEGNYMVQ ENKQEVMDGV DSFEGHDEVQ 600 CVKFSCSIPT VTGRGFIEIE DHGFSSSFFP FLVAEEDVCS EIRMLEGVLE FTETDADFEE 660 IEKMEAKNQA MNFVHEIGWL LHRIQLKSRL GHLDPSMDLF PLRRFKWLME FSMDHEWCAV 720 VRKLLNVLHN GIVGTGEHSS LNVALAEMGL LHRAVRRNSR SLVELLLRYV PEKFGSEDEA 780 LFGGSHGSIL FRPDVTGPAG LTPLHIAAGK DGSEDVLDTL TEDPGMVGIE AWKNARDSTG 840 FTPEDYARLR GHYTYIHLVQ RKINKRQAVG GHVVLDIPSN LSDSNITEKQ NEGLTSSLEI 900 GQTASRPIQR NCKLCSKKVV YGIASRSQLY RPAMLSMVAI AAVCVCVALL FKSCPEVLYV 960 FRPFRWEMLD YGTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 4e-30 | 149 | 223 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 96 | 100 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
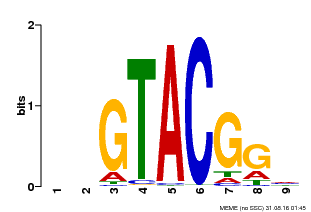
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0014s0380.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024450080.1 | 0.0 | squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2I4KKA4 | 0.0 | A0A2I4KKA4_POPTR; SBP family protein | ||||
| STRING | POPTR_0002s18970.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0014s0380.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



