 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0014s0380.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1003aa MW: 111481 Da PI: 6.403 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.6 | 1.3e-41 | 149 | 226 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C +dls+ak+yhrrhkvCe+hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
SapurV1A.0014s0380.1.p 149 VCQVEDCGVDLSNAKDYHRRHKVCEMHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 226
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.6E-33 | 145 | 211 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.308 | 147 | 224 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 5.62E-39 | 148 | 229 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.7E-30 | 150 | 223 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 2.7E-7 | 300 | 317 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.7E-7 | 403 | 413 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.21E-7 | 777 | 886 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.7E-7 | 779 | 885 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.00E-8 | 779 | 886 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1003 aa Download sequence Send to blast |
METIFGGEAR HFYAMGSTDK REVGKRGLEW NLNDWKWDGD LFIASPLNPV PSTNVSRQLF 60 PLGVGTGVPA TGNSYSSSSF SDEVNLGVEK GKRELEKRRR VVVIDDDNLN DQETGGLNLK 120 LGGERDVGNW EGSSGKKTKL VGGGLSRAVC QVEDCGVDLS NAKDYHRRHK VCEMHSKASK 180 ALVGNVMQRF CQQCSRFHVL QEFDEGKRSC RRRLAGHNKR RRKTNPDTVG NGSSMNDDQT 240 SGYLLIRLLR ILSNMHSNRS NETTDQDLLT HLLRSLASHS VEHGGRNLFG LLQEPRDLST 300 SFGNSEVVST LLSNGEGPSN LKLHLTVPVS GMPQQVMRMH DANGANIQTT FSLKPSIPNN 360 FATYSEVRES TAVQVKMNNF DLNDIYVDSD DGTEDIERSP APVNARTSSL DCPSWVQKDS 420 HQSSPPQTSR NSDSASAQSP SSSGGEAQCR TDRIVFKLFG KEPNDFPLVL RAQILDWLSH 480 SPTDIESYIR PGCIILTIYL HQAEAAWEEL CCGLGSSLSR LLDVSEDTFW RTGWVYIRVQ 540 HQIAFVYNGQ VVVDTSLPLT SNNCSKILSV KPIAITATER TEFLIKGINL SRPATKLLCA 600 VEGNYMVQEN KQEVMDGVDS FEGHDEVQCV KFSCSIPTVT GRGFIEIEDH GFSSSFFPFL 660 VAEEDVCSEI RMLEGVLEFT ETDADFEEIE KMEAKNQAMN FVHEIGWLLH RIQLKSRLGH 720 LDPSMDLFPL RRFKWLMEFS MDHEWCAVVR KLLNVLHNGI VGTGEHSSLN VALAEMGLLH 780 RAVRRNSRSL VELLLRYVPE KFGSEDEALF GGSHGSILFR PDVTGPAGLT PLHIAAGKDG 840 SEDVLDTLTE DPGMVGIEAW KNARDSTGFT PEDYARLRGH YTYIHLVQRK INKRQAVGGH 900 VVLDIPSNLS DSNITEKQNE GLTSSLEIGQ TASRPIQRNC KLCSKKVVYG IASRSQLYRP 960 AMLSMVAIAA VCVCVALLFK SCPEVLYVFR PFRWEMLDYG TS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-30 | 149 | 223 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 96 | 100 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
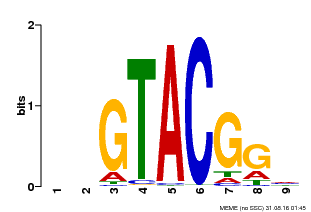
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0014s0380.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024450078.1 | 0.0 | squamosa promoter-binding-like protein 1 isoform X1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | B9GSZ5 | 0.0 | B9GSZ5_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0002s18970.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0014s0380.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



