 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo3G0024000 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 376aa MW: 41595.8 Da PI: 5.5027 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 30.4 | 6.8e-10 | 307 | 341 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
+ +yp++ee+++LA+ +gL+++q+ +WF N+R ++
Spipo3G0024000 307 RWPYPTEEEKTKLAEVTGLDQKQINNWFINQRKRH 341
569*****************************885 PP
| |||||||
| 2 | ELK | 34.5 | 4.3e-12 | 261 | 282 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++LlrKYs yL+sLk+EF+
Spipo3G0024000 261 ELKEMLLRKYSAYLSSLKKEFL 282
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 0.013 | 67 | 116 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 6.3E-8 | 69 | 92 | IPR005540 | KNOX1 |
| SMART | SM01255 | 16 | 120 | 152 | IPR005540 | KNOX1 |
| SMART | SM01256 | 6.1E-24 | 155 | 206 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 2.7E-23 | 159 | 205 | IPR005541 | KNOX2 |
| SMART | SM01188 | 1300 | 164 | 184 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 10.72 | 261 | 281 | IPR005539 | ELK domain |
| Pfam | PF03789 | 3.2E-9 | 261 | 282 | IPR005539 | ELK domain |
| SMART | SM01188 | 3.4E-6 | 261 | 282 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 13.135 | 281 | 344 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.71E-19 | 283 | 349 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 4.5E-14 | 283 | 348 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-26 | 286 | 346 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.67E-13 | 293 | 345 | No hit | No description |
| Pfam | PF05920 | 2.4E-17 | 301 | 340 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 319 | 342 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 376 aa Download sequence Send to blast |
MEDLYSIHPG IAHSDDAAVG TMSDSDSTLF HTDASSAAAL HRKVHGGERV PRELLGASTS 60 SASDLSDVVK TQISRHPRFP ALVSAYIDCR KVKYSASLFA KSIVPPLDCN LRLFSPGYYY 120 TCLLCRFSSW KQVGAPPEVA SLLEEIRRGN YPIAACSEVG TDPELDEFME SYCEVLHRFK 180 EELSKPFDEA TSFLSSIEVQ LSDLCRITAP APAAASQSTT TTATATSAGE GGGSVEEEEP 240 SSEEAEATES QETAPRMGDQ ELKEMLLRKY SAYLSSLKKE FLKKRKKGKL PKDARMELMA 300 WWKTHYRWPY PTEEEKTKLA EVTGLDQKQI NNWFINQRKR HWKPSEDMRS ALLEGVAGGS 360 GGNAVLYMDT RTVDP* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 276 | 285 | LKKEFLKKRK |
| 2 | 282 | 286 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
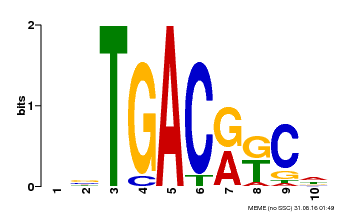
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC254547 | 0.0 | AC254547.1 Spirodela polyrhiza clone GNAS837-G21, complete sequence. | |||
| GenBank | AC254559 | 0.0 | AC254559.1 Spirodela polyrhiza clone GNAS837-J05, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008781110.1 | 1e-118 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 9e-97 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | A0A1D1Z1V5 | 1e-125 | A0A1D1Z1V5_9ARAE; Homeobox protein knotted-1-like 1 | ||||
| STRING | XP_008781110.1 | 1e-118 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1698 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G23380.2 | 2e-63 | KNOTTED1-like homeobox gene 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo3G0024000 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



