 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo27G0017800 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 451aa MW: 49127.7 Da PI: 5.958 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 90.6 | 1.4e-28 | 227 | 290 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGs........ekAtPktilelmkvkgLtlehvkSHLQkYRla 56
kpr++Wtp+LH+rFv+a++ LGGs ++AtPk+i+elm+v+gLt+++vkSHLQkYRl+
Spipo27G0017800 227 KPRRCWTPDLHRRFVNALQILGGSqgefpttlHVATPKQIRELMNVDGLTNDEVKSHLQKYRLH 290
79*********************84444444459****************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.79E-15 | 224 | 293 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-27 | 224 | 293 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.8E-23 | 227 | 290 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.6E-6 | 229 | 288 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 451 aa Download sequence Send to blast |
MERESSQKAW MGSPSELTLD FRPHGHSAAL RAIADQSDQT QNLEDFLSRL EKERLKIEAF 60 KRELPLCMQL LNNAMEAYRQ QLDMHHTNRS PRPVLEEFIP LKNPSVAAEA PEKSAAADDV 120 LSEKTSWMTT AQLWGTAEGA AAVQRTVTPP EATDDKLRTG GAFLPFSKER DGRVRGGLPE 180 LALAPMEKED EKRPYSENGG KGGAAADHAK AAAADGGHAT AVQTQRKPRR CWTPDLHRRF 240 VNALQILGGS QGEFPTTLHV ATPKQIRELM NVDGLTNDEV KSHLQKYRLH TRRPISAPQA 300 SAAAAPPQLV VLGGIWVPPE YTASSTTAKA GQLFYGGHPA AAHYCAAAVQ QEFYSPPLQP 360 PHYHHGTASA ARAALAAPHE MKRESPESEM RRSPGSTHST EEDEEVEGDD DDDDVEEGRP 420 SEAAGGKEVD DKEAAPLRED GKLSGVRLKF * |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
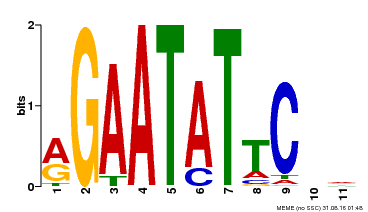
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008792133.1 | 1e-127 | transcription factor NIGTH1 | ||||
| TrEMBL | A0A1D1XR82 | 1e-130 | A0A1D1XR82_9ARAE; Two-component response regulator ARR18 (Fragment) | ||||
| STRING | XP_008792133.1 | 1e-126 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9773 | 33 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 5e-84 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo27G0017800 |



