 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo1G0061100 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 792aa MW: 87538.1 Da PI: 5.7041 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 78.3 | 7.9e-25 | 145 | 246 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++p++ ae+ ++++++ ++l+++d++ +sW++++iyr++++r++lt+GW+ Fv a++Lk+gD+v+F ++++
Spipo1G0061100 145 FCKTLTASDTSTHGGFSVPRRAAEKLfpqldySMQPPN-QELIVQDLHDNSWTFRHIYRGQPKRHLLTSGWSVFVGAKRLKAGDSVLFI--RDEK 236
99*********************999*****9555555.49************************************************..4567 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+l+++++r+
Spipo1G0061100 237 SQLLLGIRRA 246
7789999997 PP
| |||||||
| 2 | Auxin_resp | 119.7 | 2.3e-39 | 270 | 352 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaa+++s+F+v+YnPra++seFvv+++k++ka ++++svG+Rf m+feteds +rr++Gtvvg+sd+dp+rWpnSkWr+L+
Spipo1G0061100 270 AAHAAANRSPFTVYYNPRACPSEFVVPIAKYHKAAYNQISVGTRFGMMFETEDSGKRRYMGTVVGLSDCDPLRWPNSKWRNLQ 352
79*******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.28E-42 | 133 | 273 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-41 | 138 | 256 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.50E-22 | 144 | 245 | No hit | No description |
| PROSITE profile | PS50863 | 13.239 | 145 | 247 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.0E-22 | 145 | 246 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.7E-25 | 145 | 247 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 5.3E-35 | 270 | 352 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 24.492 | 686 | 770 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 1.57E-8 | 688 | 765 | No hit | No description |
| Pfam | PF02309 | 6.3E-8 | 730 | 774 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 792 aa Download sequence Send to blast |
MSSVEEKARP WDAGNARTLI EEMKLLGEFK SQPGTRKVIN SELWHACAGP LISLPQPGSL 60 VYYFPQGHSE QVTASTKRIV NSHIPNYPSL PYQMMCQVHS VTLHADKDTD EIYAQMTLQP 120 VNSENDVFPI ADFGLTKSKH PSEFFCKTLT ASDTSTHGGF SVPRRAAEKL FPQLDYSMQP 180 PNQELIVQDL HDNSWTFRHI YRGQPKRHLL TSGWSVFVGA KRLKAGDSVL FIRDEKSQLL 240 LGIRRANRQQ TVLPSVLSAD SMHLGVFAAA AHAAANRSPF TVYYNPRACP SEFVVPIAKY 300 HKAAYNQISV GTRFGMMFET EDSGKRRYMG TVVGLSDCDP LRWPNSKWRN LQVEWDEHSS 360 VERPDRVSMW EIETPESLFV VPTLASNLKR QLSPGIIGNY YEENSASHSN PEDLSTQEAP 420 YLAPAPTLMG DLQHMQDQSR LGAISSVDVN NFHSGHLASD YLDNAWWLLG STCHHSALAG 480 LNPPELLSSS LKQGSLLLCP SGNSTTNSAD ISNVLDPEIS AATDGLQFSL LSETNHRILQ 540 SLGHGFLNHN QATQDAKLLQ SISNPYELRD FADGNMNQNG AFSNLPLKSH SGDDRADPYH 600 SAGAECGLDA GKLCCLLMPS EGFHSNCNSC QDVQSQVTTA SLTDCRTVSF QEFPDCSGGT 660 SSSNLEFDDL YHLNTGSWKQ VPPPLRTFTK VQKAGSVGRS IDATRFRNYD ELKNAIACMF 720 GLEGQLDDPR GSGWKLVYVD YENDVLLVGD DPWEEFITCV KCIRILSPSE VQQMSQEGLQ 780 LMNMVTGAGE L* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 18 | 377 | 32 | 392 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional activator. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Mediates embryo axis formation and vascular tissues differentiation. Functionally redundant with ARF7. May be necessary to counteract AMP1 activity. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:14973283, ECO:0000269|PubMed:17553903}. | |||||
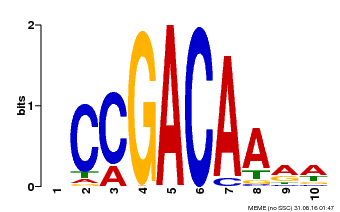
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024166618.1 | 0.0 | auxin response factor 5 isoform X2 | ||||
| Swissprot | P93024 | 0.0 | ARFE_ARATH; Auxin response factor 5 | ||||
| Swissprot | Q01K26 | 0.0 | ARFK_ORYSI; Auxin response factor 11 | ||||
| Swissprot | Q8S983 | 0.0 | ARFK_ORYSJ; Auxin response factor 11 | ||||
| TrEMBL | A0A1D1YNB9 | 0.0 | A0A1D1YNB9_9ARAE; Auxin response factor | ||||
| STRING | Migut.C00816.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP8673 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo1G0061100 |



