 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sopim01g095460.0.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum; Lycopersicon
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 419aa MW: 44616.2 Da PI: 5.8086 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 69.7 | 4.5e-22 | 275 | 337 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkre+rkq+NRe+ArrsR+RK+ae eeL+ +v++L++eN +L++e+++l ++++klk ++
Sopim01g095460.0.1 275 ERELKREKRKQSNRESARRSRLRKQAEAEELAIRVQALTGENLTLRSEINKLMENSEKLKLDN 337
89**********************************************************887 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 1.1E-37 | 1 | 92 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 1.0E-36 | 130 | 255 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 5.7E-16 | 269 | 334 | No hit | No description |
| Pfam | PF00170 | 6.1E-21 | 275 | 337 | IPR004827 | Basic-leucine zipper domain |
| SMART | SM00338 | 1.3E-18 | 275 | 339 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 12.392 | 277 | 340 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 5.13E-10 | 278 | 334 | No hit | No description |
| CDD | cd14702 | 5.32E-20 | 280 | 328 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 282 | 297 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 419 aa Download sequence Send to blast |
MGNSEDGKSC KLEKPSSPAA DQSNLHVYPD WAAMQAYYGP RVAVPAYFNS AVAPGHTPHP 60 YMWGPQPMVP PYGTPYAAIY AHGGVYAHPG VPIGSHPPGH VMATSPVVSQ AMDGASLSLD 120 ASAKSSGNSD RGLMSKLKEF EGLAMSLGNG STDNVEGGTD NGHSQSGETE GSTDGSDTNG 180 AGVSERTKKR SCETTPDNSG DDKSHSQRFQ PTGEVNDDAE KAIVVAAKMT GTVLSPCMTT 240 LEMRNPASAH MKSSPTNGGS PLSPALPNET WLQNERELKR EKRKQSNRES ARRSRLRKQA 300 EAEELAIRVQ ALTGENLTLR SEINKLMENS EKLKLDNATL MERLKNEQLG QTEEVSLGKI 360 DDKRLQPVGT VNLLARVNNS GSSDTTNEDG EVYENNSSGA KLHQLLDTSP RTDAVAAG* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 291 | 297 | RRSRLRK |
| 2 | 291 | 298 | RRSRLRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-activator of a beta-glucuronidase (GUS) reporter gene. Binds to a G-box-related element, (5'-GCAACGTGGC-3'). Also binds to the HEX-motif of wheat histone H3 promoter. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
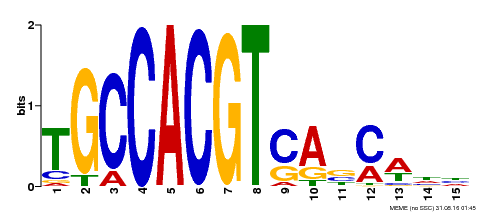
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013341 | 0.0 | BT013341.1 Lycopersicon esculentum clone 135142F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004229974.1 | 0.0 | transcriptional activator TAF-1 isoform X4 | ||||
| Swissprot | Q99142 | 1e-147 | TAF1_TOBAC; Transcriptional activator TAF-1 (Fragment) | ||||
| TrEMBL | A0A3Q7ELE7 | 0.0 | A0A3Q7ELE7_SOLLC; Uncharacterized protein | ||||
| STRING | Solyc01g095460.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3113 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 1e-115 | G-box binding factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



