 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sopen01g030050.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum; Lycopersicon
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 983aa MW: 109159 Da PI: 6.1987 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.9 | 9e-41 | 134 | 211 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C +dls+ak+yhrrhkvCe+hska+ +lv +++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+q+
Sopen01g030050.1 134 VCQVDDCGTDLSKAKDYHRRHKVCEMHSKASRALVGNVMQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNKRRRKTQS 211
6**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.0E-32 | 130 | 196 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.086 | 132 | 209 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 6.8E-38 | 133 | 214 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.6E-30 | 135 | 208 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 2.63E-7 | 754 | 867 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.9E-5 | 756 | 868 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.15E-6 | 757 | 867 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 983 aa Download sequence Send to blast |
MGGTDLRGLG KRSLEWDLTD WKWDGDLFIA TPLQQNPSNY QSRQFFPVET GNLASSNSSS 60 SCSDEVNHGM EQQRRELEKR RRVIVVDEDD SGPLSLKLGG QGEPAADAGR EMSNWDGAAG 120 KRTKLAAPAA TRAVCQVDDC GTDLSKAKDY HRRHKVCEMH SKASRALVGN VMQRFCQQCS 180 RFHALQEFDE GKRSCRRRLA GHNKRRRKTQ SETVANNNSL NDGQTSGYSL MSLLKILSNM 240 HSNGANHTED QDLLSHLLRS LASQGPTNGD KSLSGLLQES SNLLNNRSIL RNPEIASLIS 300 NGSQAPPRPK ERQFTNSAAE MPQKRLEDAR TASSQSPGIL FPIQSNSQAY TPGRESTTGR 360 SKLIDFDLND AYVDSDDCGD DIDRSPVPEC PSWLQQDSHQ SSPPQTSGNS DSASAQSPSS 420 SSGDNQNRTD RIVFKLFGKG PSDFPFVVRA QILDWLSHSP TEIESYIRPG CVVLTIYLRL 480 PESAWEELSY DLSSSLSRLL DVHGGDSFWT KGWIYIRVQN QIAFVCDGQV LLDMSLPCVS 540 NDDSTLLSVR PIAVPVSDRV QFLVKGYNLT KPSTRLLCSL EGNYLDPEAD NEVEKQVAGG 600 DKDDKLQSLN FTCSIPAVGG RGFIEVEDHG VSNSFFPFII AEEDVCSEIR MLESDLELTS 660 LDYVKGQTNN IEARNQAMDF IHELGWLLHR NNLRARLEHF GPNAVLHPLK RFKWLVEFSV 720 DHEWCAVVKK LLNILLDGTV GGDDSSLKYA LTEMGLLHKA VRRNSRPLVE LLLTYTPTNV 780 ADELCCEYQS LVGVGGQFLF RPDCVGPGGL TPLHIAAGID GYEDVLDALT DDPGKVAIEA 840 WKNTRDSTGF TPEDYARLRG HYSYIHLVQR KISKKANSGH IVVDIPRVPS VVENSNQKDE 900 VCATTSLEIS MTERKAIPRP CRLCDRKLAY GSRSRSLLYR PAMFSMVAMA AVCVCVALLF 960 RGSPEVLYIF RPFRWEMVDF GTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 7e-31 | 134 | 208 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 78 | 82 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
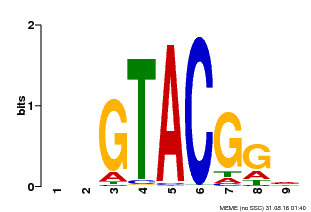
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975440 | 0.0 | HG975440.1 Solanum pennellii chromosome ch01, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015055592.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A3Q7EGS2 | 0.0 | A0A3Q7EGS2_SOLLC; Uncharacterized protein | ||||
| STRING | Solyc01g068100.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



