 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_165590_jids.t1 | ||||||||
| Common Name | SOVF_165590 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 843aa MW: 91396.3 Da PI: 6.2449 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.7 | 3.1e-21 | 145 | 200 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe++F+++++p++++r eL+kkl L++rqVk+WFqNrR+++k
Sp_165590_jids.t1 145 KKRYHRHTPQQIQELEAVFKECPHPDEKQRLELSKKLCLETRQVKFWFQNRRTQMK 200
688999***********************************************999 PP
| |||||||
| 2 | START | 195.5 | 2.4e-61 | 358 | 584 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE......EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S.... CS
START 1 elaeeaaqelvkkalaeepgWvkss......esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla.... 77
ela++a++elvk+a+++ep+W +s e++n +e++++f + + + ea r+sg+v+ ++ lve+l d + +W e+++
Sp_165590_jids.t1 358 ELALAAMDELVKLAQSDEPLWLRSMdaaggrEILNHEEYMRSFNPCIGmkpngFVPEATRDSGMVIINSLALVETLTDAN-RWAEMFPgmia 448
5899***************************************9988899999999************************.*********** PP
EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEE CS
START 78 kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghs 162
+a+t++vissg galqlm aelq+lsplvp R + f+R+++q+ +g+w++vdvS+d ++ + s+ +v +++lpSg+++++++ng+s
Sp_165590_jids.t1 449 RATTTDVISSGmggtrnGALQLMHAELQVLSPLVPvRVVNFLRFCKQHAEGVWAVVDVSIDCIRESSGSPTYVNCRRLPSGCVVQDMPNGYS 540
*****************************************************************999************************ PP
EEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 163 kvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
kvtwveh++++++++h+l+rslv++g+ +ga++w tlqrqce+
Sp_165590_jids.t1 541 KVTWVEHAEYDESSVHQLFRSLVSTGMGFGAQRWITTLQRQCEC 584
******************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.63E-20 | 131 | 202 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.232 | 142 | 202 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.7E-18 | 143 | 206 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-21 | 143 | 210 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.84E-19 | 144 | 202 | No hit | No description |
| Pfam | PF00046 | 9.4E-19 | 145 | 200 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 177 | 200 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.348 | 349 | 587 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.92E-32 | 351 | 584 | No hit | No description |
| CDD | cd08875 | 2.97E-121 | 353 | 583 | No hit | No description |
| SMART | SM00234 | 1.7E-45 | 358 | 584 | IPR002913 | START domain |
| Pfam | PF01852 | 1.1E-53 | 358 | 584 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.57E-25 | 611 | 836 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 843 aa Download sequence Send to blast |
MNFGGFLDSS GAGAEAGGGG GGGAKLVTDI SYNNNMPPGA ITHHQQQQLQ QQQQQRLVGP 60 TSINNKSFFN TPGLSLALQT GMEGHGEVTR VTEQNYDGNS NGGGGGVSVG VRRSREDEHE 120 SRSGSDNLDG ASGDEQDIAD LPPRKKRYHR HTPQQIQELE AVFKECPHPD EKQRLELSKK 180 LCLETRQVKF WFQNRRTQMK TQLERHENSI LRQENDKLRA ENMSIREAMR NPICTNCGGQ 240 AMIGDISMEE QHLRIENARL KDELDRVCSL ASKFLGRPVS SLSTLSGAIS APPMPNSSLE 300 LAVGTNGFGP LTTVTTSSMP LGDYGSGISN AMPGVSPTKS TANVTGMERS FERSMYLELA 360 LAAMDELVKL AQSDEPLWLR SMDAAGGREI LNHEEYMRSF NPCIGMKPNG FVPEATRDSG 420 MVIINSLALV ETLTDANRWA EMFPGMIARA TTTDVISSGM GGTRNGALQL MHAELQVLSP 480 LVPVRVVNFL RFCKQHAEGV WAVVDVSIDC IRESSGSPTY VNCRRLPSGC VVQDMPNGYS 540 KVTWVEHAEY DESSVHQLFR SLVSTGMGFG AQRWITTLQR QCECLAILMS SNIPSGDHTA 600 ITASGRRSML KLAQRMTNNF CAGVCASSVH KWNKLNIGNV DEDVRVMTRK SIDDPGEPSG 660 IVLSAATSVW LPVSQQRLFD FLRDERLRSE WDILSNGGPM QEMAHIAKGQ DQGNCVSLLR 720 AGAMNPNQSS MLILQETCTD SSGSLVVYAP VDIPAMHVVM NGGDSAYVAL LPSGFAIVPD 780 GPGSRGTNNG SPQRGGGSLL TVAFQILVNS LPTAKLTVES VETVNNLISC TVQKIKAALN 840 CES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
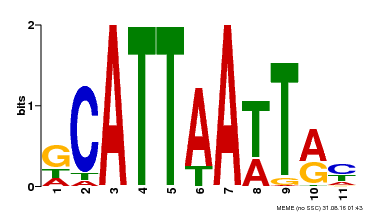
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021849370.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2-like | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A0K9QLI3 | 0.0 | A0A0K9QLI3_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010671720.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



