 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_146430_eotc.t1 | ||||||||
| Common Name | SOVF_146430 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 329aa MW: 36559.3 Da PI: 9.009 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43 | 1.1e-13 | 63 | 107 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
Sp_146430_eotc.t1 63 RENWTEQEHDKFLEALQLFDRD-WKKIEAFIG-SKTVIQIRSHAQKY 107
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.17E-16 | 57 | 113 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.95 | 58 | 112 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.9E-7 | 61 | 113 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.9E-18 | 61 | 110 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 6.0E-11 | 62 | 110 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.2E-11 | 63 | 107 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.68E-8 | 65 | 108 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 329 aa Download sequence Send to blast |
MVSVNPNPLQ PQGYNLEFNS DMPLTGVNSL PPVNSPVSSG YVVPATEDHC KKVRKPYTIT 60 KSRENWTEQE HDKFLEALQL FDRDWKKIEA FIGSKTVIQI RSHAQKYFLK VQKNGSNERV 120 PPPRPKRKAA HPYPQKASKI ASTTSQSTGQ YQESPVCALE PGYVYHHDPS SVLTNQSGSG 180 STSPWTFGSM PVVNSANMRR DDAALAMSMT PTGCSSSSNE SGSTWPTVGT MDQKHTKAVP 240 DFAQVYSFIG SVFDPNARDH LQRLKMMDPI NVETVLLLMK NLSVNLLSPE FEEYRKLFSS 300 YDVESGKTRS CSHLLPAPNN SKTVIPAVY |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
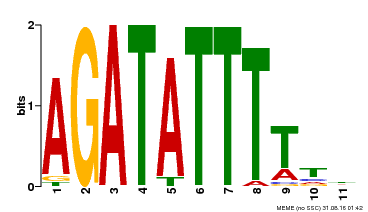
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00423 | DAP | Transfer from AT4G01280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FJ752586 | 1e-51 | FJ752586.1 Beta vulgaris clone BAC SBI-153H13, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021850430.1 | 0.0 | protein REVEILLE 5 isoform X1 | ||||
| Swissprot | C0SVG5 | 6e-97 | RVE5_ARATH; Protein REVEILLE 5 | ||||
| TrEMBL | A0A0K9QSI3 | 0.0 | A0A0K9QSI3_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010669883.1 | 1e-170 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01280.1 | 3e-72 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



