 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_121730_ewts.t1 | ||||||||
| Common Name | SOVF_121730 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 695aa MW: 76666.4 Da PI: 6.6397 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.1 | 3.5e-23 | 120 | 221 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f k+lt+sd+++ g +++p+ +ae++ ++ + + +d +g+ W++++iyr++++r++lt+GW+ Fv+a++L +gD++vF
Sp_121730_ewts.t1 120 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--NILAKDVHGEPWKFRHIYRGTPRRHLLTTGWSTFVNAKKLIAGDSIVFL-- 207
89****************************997656666..69999********************************************.. PP
SSSEE..EEEEE-S CS
B3 86 grsefelvvkvfrk 99
+ +++ l+v+++r+
Sp_121730_ewts.t1 208 RAENGDLCVGIRRA 221
4488999****997 PP
| |||||||
| 2 | Auxin_resp | 107.7 | 1.3e-35 | 289 | 370 | 3 | 83 |
Auxin_resp 3 haastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
+ as++ +FevvY+Pra+t+eFvvk+ +v++al++ ++ GmRfkm fetedss++++ +Gt+++v+ +dp++W++S+Wr+L+
Sp_121730_ewts.t1 289 TYASKGHPFEVVYYPRAGTPEFVVKAGHVKAALQICWCSGMRFKMPFETEDSSRISWfMGTISSVQVADPMHWSDSPWRLLQ 370
67899***************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 7.06E-29 | 110 | 223 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.4E-36 | 114 | 223 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.14E-21 | 119 | 220 | No hit | No description |
| Pfam | PF02362 | 1.8E-21 | 120 | 221 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.042 | 120 | 222 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.4E-21 | 120 | 222 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 8.0E-31 | 289 | 370 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 18.225 | 609 | 692 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 695 aa Download sequence Send to blast |
MITFMDPKEK IKEADNCLDS QLWHACAGGM VQMPPVSSKV YYFVQGHIEH ANGNVDFRSF 60 PRIPSYVLCR VEDIRYLADQ ETDEVYAKIR LSPVTNEVDY SDDDIGVHGT EVQENKPASF 120 AKTLTQSDAN NGGGFSVPRY CAETIFPRLD YSADPPVQNI LAKDVHGEPW KFRHIYRGTP 180 RRHLLTTGWS TFVNAKKLIA GDSIVFLRAE NGDLCVGIRR AKRGLGGGLD STSGWNPGST 240 VLNYGAFSAF LREDENRIGR NGNGVGLNSS GNPSAKGKVK LESVLESATY ASKGHPFEVV 300 YYPRAGTPEF VVKAGHVKAA LQICWCSGMR FKMPFETEDS SRISWFMGTI SSVQVADPMH 360 WSDSPWRLLQ VTWDEPDLLQ NVKRVSPWLV ELVSNMPTIH LSPFSPPRKK FRLPHHPDFP 420 MDGQLPVPTF PGNFLGPSNP FGCLPDNSPA GMQGARHAQF GLSLSDLHMS KLQSGLFPGG 480 FPALDTTNTL PNRPSSSLSY HKPGSNENIS CLLTMGNTNQ SVKKSDVTKP HHFVLFGKPI 540 LTEQQMSLSR SVDTVSPVPT GNSSSEGNVD KAGNVSDGSG SGPNQDRSSM ERLPWLKDNR 600 PELDLELETG HCKVFMESED VGRTLELTLL NSYEELCTRL ANMFSIETSD MLNRVVYRDS 660 AGSSKQIGDE PFSEFAKRAK RLTILMDSSS DNTGA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 1e-81 | 18 | 391 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
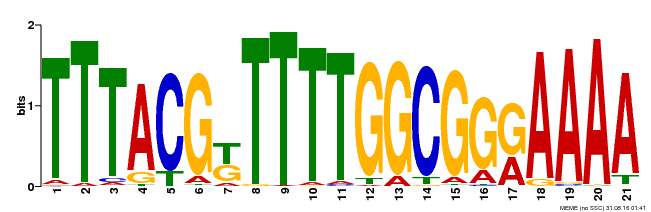
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021863267.1 | 0.0 | auxin response factor 18-like | ||||
| Refseq | XP_021863268.1 | 0.0 | auxin response factor 18-like | ||||
| Refseq | XP_021863269.1 | 0.0 | auxin response factor 18-like | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A0K9R0A2 | 0.0 | A0A0K9R0A2_SPIOL; Auxin response factor | ||||
| STRING | XP_010684071.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



