 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_041720_iowe.t2 | ||||||||
| Common Name | SOVF_041720 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 686aa MW: 75338.8 Da PI: 6.2454 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 81.8 | 7.9e-26 | 213 | 265 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++ ++Pk+ilelm+v+gLt+e+v+SHLQkYRl
Sp_041720_iowe.t2 213 KPRVVWSVELHQQFVAAVNQL-GID-TVPKKILELMNVPGLTRENVASHLQKYRL 265
79*******************.888.8***************************8 PP
| |||||||
| 2 | Response_reg | 76.8 | 7.7e-26 | 37 | 145 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedaleal 90
vl+vdD+p+ + +l+++l+ +y ev+ + +e al ll+ek+ +D+++ D+ mp+mdG++ll++ e +lp+i+++a ++++ + + +
Sp_041720_iowe.t2 37 VLVVDDDPTCLTILEKMLRTCRY-EVTKTNRAEYALSLLREKKngFDIVISDVHMPDMDGFKLLEHVGLEM-DLPVIMMSADDSKQVVMKGV 126
89*********************.***************999999**********************6644.8******************* PP
HTTESEEEESS--HHHHHH CS
Response_reg 91 kaGakdflsKpfdpeelvk 109
Ga d+l Kp+ +e+l +
Sp_041720_iowe.t2 127 THGACDYLIKPVRIEALKN 145
***************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 4.1E-166 | 12 | 656 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 3.9E-40 | 34 | 158 | No hit | No description |
| SuperFamily | SSF52172 | 6.83E-34 | 34 | 156 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 2.5E-27 | 35 | 147 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.096 | 36 | 151 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 5.6E-23 | 37 | 145 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 8.99E-25 | 38 | 150 | No hit | No description |
| PROSITE profile | PS51294 | 10.283 | 210 | 268 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.91E-18 | 210 | 269 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-27 | 212 | 270 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-22 | 213 | 265 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.6E-8 | 215 | 264 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0031537 | Biological Process | regulation of anthocyanin metabolic process | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0080036 | Biological Process | regulation of cytokinin-activated signaling pathway | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 686 aa Download sequence Send to blast |
MNLGGGLMGS MSMPSSTASR KSVDSVTGDV FPVGLRVLVV DDDPTCLTIL EKMLRTCRYE 60 VTKTNRAEYA LSLLREKKNG FDIVISDVHM PDMDGFKLLE HVGLEMDLPV IMMSADDSKQ 120 VVMKGVTHGA CDYLIKPVRI EALKNIWQHV VRKKKYDYKD VEQSGSWDEG DRQLKQDDAV 180 SSPANDGSWK NSKRKSGEDE DTDEKDDNAT LKKPRVVWSV ELHQQFVAAV NQLGIDTVPK 240 KILELMNVPG LTRENVASHL QKYRLYLRRL SGVSQHQGGL NNSFMPQDPT FSTISSLGGI 300 DLQTLAATGQ FSAQTLAAYT RLPPSIKPGM SMPFVDQRSL FSFENSKLRY GEGQQQQQQQ 360 QQQQQAQINN VNKQMNLLHG IPTTMEPKQL VGLNQSAQTL GSMSMSMSMQ GNASSSHQSS 420 SLLMQQMVPQ QSRGQISNEC VSSQVSRIQP SIIGQQLQPN GNATSVLSRN GMGGYDTVSQ 480 AGPVVDFSVN QISELSGNSF PLGSTPGITS IASKGYNQEE ISSDIKVTRG FVGSYDMFND 540 HLHHKSQEWP LQNPNMFAGS SQHVVASMQS GLNVAPSVMV NQSYVPAQKI EQNGRAIAGK 600 PMFSAGLENH QHMGMQNVNQ NFNSVLVNNS ARVKAEAVSD VVNLGDNLFP ERYSQEDLMA 660 VLKQQDGIGS GGEFDFDGYT LDNIPV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 7e-20 | 212 | 271 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
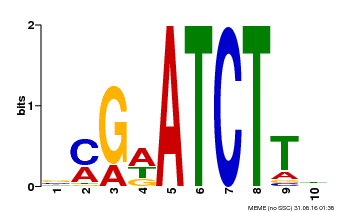
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021848392.1 | 0.0 | two-component response regulator ARR2-like isoform X3 | ||||
| Swissprot | Q9ZWJ9 | 1e-177 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A0K9RPY6 | 0.0 | A0A0K9RPY6_SPIOL; Two-component response regulator | ||||
| STRING | XP_010670914.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-173 | response regulator 2 | ||||



