 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_038230_pxgw.t1 | ||||||||
| Common Name | SOVF_038230 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 419aa MW: 46904.6 Da PI: 6.6777 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 74.5 | 3.2e-23 | 47 | 148 | 3 | 66 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt............................. 65
g+kdrhsk++T +g+RdRR+Rls+++a++++dLqd+LG+ ++sk i+WLl k i++l+ +
Sp_038230_pxgw.t1 47 GGKDRHSKVCTIKGLRDRRIRLSVPTAIQLYDLQDRLGLSQPSKVIDWLLDTTKLDIDKLPPLppipggfnpsqgygmpptpffdasllpsr 138
78************************************************************988888777777776666666655555554 PP
TCP 66 .........s 66
+
Sp_038230_pxgw.t1 139 kecgininsT 148
4444444440 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51369 | 28.364 | 48 | 106 | IPR017887 | Transcription factor TCP subgroup |
| Pfam | PF03634 | 5.7E-25 | 48 | 108 | IPR005333 | Transcription factor, TCP |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 419 aa Download sequence Send to blast |
MMNLNKDGPP YTTKQEEGDS SKAAAPSRQW SSGFKNPRIV RVSRTFGGKD RHSKVCTIKG 60 LRDRRIRLSV PTAIQLYDLQ DRLGLSQPSK VIDWLLDTTK LDIDKLPPLP PIPGGFNPSQ 120 GYGMPPTPFF DASLLPSRKE CGININSTMN DDVDVDVDVD VDVDVDVDEE DESSIVAKFK 180 YWGDMNAMTM MRSKQKGVDY HHEQGSSSSS QVLAQNLFPI ASSNNTTTNT NNNNNSMLPH 240 PHPHPHPHPH PHPPSFLSNN PMYNYYHLEP SNLSLSQLGS YNDHHHHQGA SQSSSTPQPP 300 SSLSLSSMSQ LFLCPSMATT SFFPSNHHAS FMNNTSHHHQ QQHQQQQVEN DLRQFNRFPL 360 SNNNNPFSSS FHNNNSSGLD MKPAFPLSMN HDPHLQQEDD EDRQGKSSAH NPWKESSGN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 3e-19 | 53 | 107 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 3e-19 | 53 | 107 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
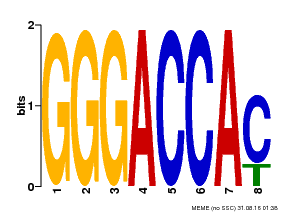
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00065 | PBM | Transfer from AT5G60970 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021856336.1 | 0.0 | transcription factor TCP17 | ||||
| Refseq | XP_021856337.1 | 0.0 | transcription factor TCP17 | ||||
| TrEMBL | A0A0K9RR59 | 0.0 | A0A0K9RR59_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010695237.1 | 1e-151 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G60970.1 | 1e-34 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 | ||||



