 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_025660_sqcn.t2 | ||||||||
| Common Name | SOVF_025660 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 371aa MW: 40911.6 Da PI: 8.8577 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 50.6 | 3.2e-16 | 195 | 253 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS-....-HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknryp....saeereeLAkklgLterqVkvWFqNrRakek 56
rk+ ++tk+q +Lee F++++++ s++++ LAk+lgL rqV vWFqNrRa+ k
Sp_025660_sqcn.t2 195 RKKLRLTKDQSAILEESFKEHNTLnplsSQKQKLALAKRLGLRPRQVEVWFQNRRARTK 253
78889**************866652233567777899********************98 PP
| |||||||
| 2 | HD-ZIP_I/II | 118.6 | 3.5e-38 | 195 | 288 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLep....erKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLr 88
+kk+rl+k+q+++LEesF+e+++L+p ++K +la++Lgl+prqv+vWFqnrRARtk+kq+E+d+e+Lkr++++l+een+rL+kev+eLr
Sp_025660_sqcn.t2 195 RKKLRLTKDQSAILEESFKEHNTLNPlssqKQKLALAKRLGLRPRQVEVWFQNRRARTKLKQTEVDCEFLKRCCEQLTEENRRLQKEVQELR 286
69**********************97666689999********************************************************9 PP
HD-ZIP_I/II 89 eel 91
+l
Sp_025660_sqcn.t2 287 -AL 288
.55 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04618 | 1.0E-17 | 5 | 157 | IPR006712 | HD-ZIP protein, N-terminal |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-17 | 177 | 253 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 7.52E-16 | 183 | 256 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.313 | 191 | 255 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.6E-12 | 193 | 259 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.1E-13 | 195 | 253 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.40E-13 | 195 | 256 | No hit | No description |
| PRINTS | PR00031 | 4.3E-5 | 226 | 235 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 230 | 253 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 4.3E-5 | 235 | 251 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.7E-11 | 255 | 289 | IPR003106 | Leucine zipper, homeobox-associated |
| SMART | SM00340 | 7.9E-27 | 255 | 298 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009641 | Biological Process | shade avoidance | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 371 aa Download sequence Send to blast |
MMVAGKEDLG LSLSLGSSNN TSNIASTSGN NNNNNHNLLT PNINQNNNIG FFMNHNNINR 60 VHPHQPHQTT SSNASPLQLN LMMMPSLACP SPSPLLQKLP WNRSTDSFPP SSDRNSDSSR 120 SFLRGIDVNR PAGGDEEDAG VSSPNSTVST VSGKRSSGER DSTTTAGAGD EIEVERASSR 180 GLSDGDDDDG GENSRKKLRL TKDQSAILEE SFKEHNTLNP LSSQKQKLAL AKRLGLRPRQ 240 VEVWFQNRRA RTKLKQTEVD CEFLKRCCEQ LTEENRRLQK EVQELRALKL SPQFYMQMTP 300 PTTLTMCPSC ERVAVPTSAT TSTTPAQPHP QPQIPDPRPN PSAHHHRTMP VNPWAAAPVP 360 IRSFEALRPR S |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 193 | 199 | SRKKLRL |
| 2 | 247 | 255 | RRARTKLKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that plays a role in auxin-mediated morphogenesis. Negatively regulates lateral root elongation. {ECO:0000269|PubMed:12492842}. | |||||
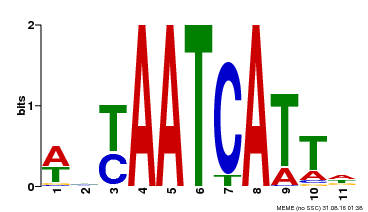
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00550 | DAP | Transfer from AT5G47370 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By indole-3-acetic acid (IAA). {ECO:0000269|PubMed:12492842}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021856175.1 | 0.0 | homeobox-leucine zipper protein HAT4-like isoform X1 | ||||
| Swissprot | P46601 | 1e-101 | HAT2_ARATH; Homeobox-leucine zipper protein HAT2 | ||||
| TrEMBL | A0A0K9RUX7 | 0.0 | A0A0K9RUX7_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010677086.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47370.1 | 3e-81 | Homeobox-leucine zipper protein 4 (HB-4) / HD-ZIP protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



