 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_025120_zkzs.t1 | ||||||||
| Common Name | SOVF_025120 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 837aa MW: 92096.6 Da PI: 6.4176 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.3 | 6.5e-19 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Sp_025120_zkzs.t1 17 KYVRYTPEQVEALERLYHECPKPSSLRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 180.1 | 1.3e-56 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..gal 91
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g++
Sp_025120_zkzs.t1 161 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPSWYRDCRAVDVLNVLPTAnsGTI 251
7899******************************************************.8888888888*********************** PP
EEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHH CS
START 92 qlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphw 179
+l +++l+a+++l+p Rdf+ +Ry+ +++g++v++++S++++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +++++
Sp_025120_zkzs.t1 252 ELLYMQLYAPTTLAPaRDFWLLRYTSVMEDGSLVVCERSLNNTQNGPSmptVQNFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPE 343
***********************************************999889*************************************** PP
HHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 180 llrslvksglaegaktwvatlqrqce 205
+lr+l++s+++ ++kt++a+l+++++
Sp_025120_zkzs.t1 344 VLRPLYESSTVIAQKTTMAALRQLRQ 369
**********************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.548 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.3E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.25E-16 | 17 | 77 | No hit | No description |
| SuperFamily | SSF46689 | 5.56E-17 | 17 | 80 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 1.6E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.65E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 26.564 | 151 | 366 | IPR002913 | START domain |
| CDD | cd08875 | 1.04E-78 | 155 | 371 | No hit | No description |
| SMART | SM00234 | 6.8E-39 | 160 | 370 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 1.4E-23 | 160 | 367 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 4.94E-38 | 160 | 372 | No hit | No description |
| Pfam | PF01852 | 4.3E-54 | 161 | 369 | IPR002913 | START domain |
| Pfam | PF08670 | 6.3E-52 | 694 | 836 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 837 aa Download sequence Send to blast |
MMAMSCKDGK GVMDNGKYVR YTPEQVEALE RLYHECPKPS SLRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSHL VYENGYFRQQ 120 TQKSGVASKD TSCESVVTSG QHHLTPQHPP RDASPAGLLS IAEETLAEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIVAISH GCTGVAARAC GLVGLEPTRV AEILKDRPSW YRDCRAVDVL 240 NVLPTANSGT IELLYMQLYA PTTLAPARDF WLLRYTSVME DGSLVVCERS LNNTQNGPSM 300 PTVQNFVRAE MLPSGYLIRP CEGGGSIIHI VDHMDLEPWS VPEVLRPLYE SSTVIAQKTT 360 MAALRQLRQI AQEVSQPNVS NWGRRPAALR ALSQRLNRGF NEALNGFTDE GWSMIGNDGI 420 DDVTVLVNSS PDKLMALNLT FANGFPAVSN TVLCAKASML LQNVPPAVLL RFLREHRSEW 480 ADNNIDAYLA AAAKISPCSL PGARIGGFGN QVILPLAHTI EHEELLEVIK LEGVVSCPED 540 AMMPRDMFLL QLCSGMDENA IGTCAELIFA PIDASFADDA PLLPSGFRII PLDSVKEASS 600 PNRTLDLASA LEVGPSANRS SSDNSGSGNN MRSVMTIAFQ FAFESHLQDN VASMARQYVR 660 SIISSVQRVA LALSPHLGSQ AGLRSPLGNP EAHTLARWIC QSYRCFLGVE LLKSSGDGSD 720 AVLKSLWHHS DAIMCCSLKA LPVFTFANQA GLDMLETTLV ALQDITLEKI LDDHGRKTLC 780 SEFPQILQQG FACLQGGICL SSMGRPVSYE RAVAWKVLNE EENAHCICFM FMNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
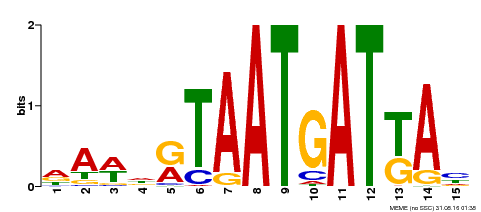
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021836126.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A0K9RV39 | 0.0 | A0A0K9RV39_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010691422.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



