 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 402040 | ||||||||
| Common Name | SELMODRAFT_402040 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Lycopodiidae; Selaginellales; Selaginellaceae; Selaginella
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 397aa MW: 44807.1 Da PI: 5.2782 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 103.9 | 9.7e-33 | 228 | 283 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++qLGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
402040 228 KARRCWSPELHRRFVNALQQLGGSQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 283
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-29 | 225 | 286 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.78E-18 | 225 | 286 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.487 | 225 | 285 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 3.5E-27 | 228 | 283 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 230 | 281 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 397 aa Download sequence Send to blast |
MNPEEEFGQA EGPDLTLGCA IAKRDSSDQQ TRRRMEDYLQ ALQQERQKIE AFKRELPCCM 60 NLLDQAIQST RGCLDESYKT EMEEENPPKL ELFLENRSGF RLGKPKSEAE DHRDFEERPS 120 TPKQNWMQLA CFPSQSPRED EEFETRKALR SSTASADSGA FAPFVRDKKW DQAELCLSSS 180 ICLSLEQKKT GFIKFQGNES NLPLVELSLQ QMPSSGICSN STLHQQRKAR RCWSPELHRR 240 FVNALQQLGG SQVATPKQIR ELMKVDGLTN DEVKSHLQKY RLHTRRPATV APGTAAQKSP 300 QVVVLGSIWV PPEYAVFDHH QAAPSMQSTP LQQSKAQDFF ANLHASPAEP MVFYEKQELL 360 KSSKEEGEVL SPRKASATTA SEEEDDGDVT EAEEQR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-14 | 228 | 281 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-14 | 228 | 281 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-14 | 228 | 281 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-14 | 228 | 281 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-14 | 228 | 281 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
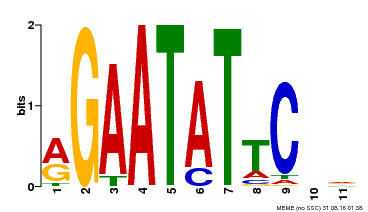
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002960068.1 | 0.0 | transcription factor HHO2 | ||||
| TrEMBL | D8QPE3 | 0.0 | D8QPE3_SELML; Uncharacterized protein | ||||
| STRING | EFJ37607 | 0.0 | (Selaginella moellendorffii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP4303 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 4e-59 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 402040 |
| Entrez Gene | 9656231 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



