 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 29692 | ||||||||
| Common Name | SELMODRAFT_113487, SELMODRAFT_29692, WRKY_2, WRKY_25 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Lycopodiidae; Selaginellales; Selaginellaceae; Selaginella
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 67aa MW: 7842.72 Da PI: 7.9299 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 89.4 | 2.9e-28 | 4 | 64 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-ST...T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsa...gCpvkkkversaedpkvveitYegeHnhe 59
dDg++WrKYGqK++ +s+fprsYYrCt++ gC+++k v++++++p++ ++tY geH ++
29692 4 DDGFTWRKYGQKDILNSKFPRSYYRCTHQkelGCQATKYVQKCEDEPSMYQVTYIGEHSCQ 64
8***************************99999*************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50811 | 20.151 | 1 | 62 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 8.3E-28 | 2 | 65 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.23E-26 | 2 | 64 | IPR003657 | WRKY domain |
| SMART | SM00774 | 4.4E-38 | 3 | 65 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 9.4E-28 | 4 | 64 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009862 | Biological Process | systemic acquired resistance, salicylic acid mediated signaling pathway | ||||
| GO:0009864 | Biological Process | induced systemic resistance, jasmonic acid mediated signaling pathway | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 67 aa Download sequence Send to blast |
APADDGFTWR KYGQKDILNS KFPRSYYRCT HQKELGCQAT KYVQKCEDEP SMYQVTYIGE 60 HSCQNAQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ir8_A | 3e-23 | 4 | 64 | 9 | 69 | OsWRKY45 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in senescence, biotic and abiotic stress responses by modulating various phytohormones signaling pathways (PubMed:14742872, PubMed:16623907, PubMed:17310369, PubMed:28576847). Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). Binds to the 5'-[CT]GACTTTT-3' motif in promoters of target genes to induce their expression (PubMed:24104863). Plays an important but not indispensable role in jasmonate and salicylic acid signaling (PubMed:18713432). Regulates positively the salicylic acid (SA)-mediated signal pathway, but negatively the jasmonic acid (JA)-mediated signal pathway, thus determining the balance between these mutually antagonistic pathways (PubMed:14742872, PubMed:16623907, PubMed:18713432, PubMed:28837631). Together with WRKY46, WRKY53 and WRKY54, prevents defense response to the necrotrophic pathogens P.carotovorum and B.cinerea, but promotes defense responses (including SA-induced pathogenesis-related (PR) genes expression) against biotrophic/hemibiotrophic SA-monitored pathogens (e.g. P.syringae, E.carotovora subsp. carotovora SCC3193 and E.cichoracearum), probably by regulating negatively the JA/ET and positively the SA signaling pathways (PubMed:28837631, PubMed:16623907, PubMed:22325892). Contributes to the suppression of jasmonic acid (MeJA)-induced expression of JA-responsive genes (e.g. PDF1.2) (PubMed:22325892, PubMed:16623907). Promotes susceptibility to JA-monitored pathogens (e.g. A.brassicicola), probably by facilitating SA-controlled suppression of JA-mediated defense. Represses the biosynthesis of the phytoalexin camalexin and indol-3-ylmethyl glucosinolate (IGS) (PubMed:16623907). Represses both SA and JA/ethylene (ET) mediated defense marker genes expression (PubMed:17310369). Negative regulator of SA biosynthesis (PubMed:28837631). Negative regulator of EDS1-dependent defense against E.amylovora (PubMed:22316300). Required for RPP4-mediated disease resistance and basal defense against H.parasitica, probably via late up-regulation (LURP) of resistance genes (e.g. CML10/CaBP22 and LURP1) (PubMed:17313163). Probably involved in defense responses toward insects (e.g. P.xylostella and B.brassicae) (PubMed:25339349). Together with WRKY54, negative regulator of developmental senescence, probably via the regulation of several senescence-associated markers genes (PubMed:17310369, PubMed:22268143). Together with WRKY46 and WRKY54, promotes brassinosteroid (BR)-regulated plant growth but prevent drought response by modulating gene expression (PubMed:28576847). In collaboration with WRKY54, prevents stomatal closure and, consequently, osmotic stress tolerance (PubMed:23815736). Regulates rhizobacterium B.cereus AR156-induced systemic resistance (ISR) to P.syringae pv. tomato DC3000 (PubMed:26433201). {ECO:0000250|UniProtKB:Q9SUP6, ECO:0000269|PubMed:14742872, ECO:0000269|PubMed:16623907, ECO:0000269|PubMed:17310369, ECO:0000269|PubMed:17313163, ECO:0000269|PubMed:18713432, ECO:0000269|PubMed:22268143, ECO:0000269|PubMed:22316300, ECO:0000269|PubMed:22325892, ECO:0000269|PubMed:23815736, ECO:0000269|PubMed:24104863, ECO:0000269|PubMed:25339349, ECO:0000269|PubMed:26433201, ECO:0000269|PubMed:28576847, ECO:0000269|PubMed:28837631, ECO:0000303|PubMed:28837631}. | |||||
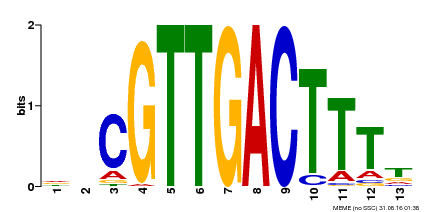
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00408 | DAP | Transfer from AT3G56400 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Regulated by MYB44 (PubMed:23067202, PubMed:23603962). Basal expression levels require the presence of endogeneous salicylic acid (SA) (PubMed:14742872, PubMed:17310369). Induced by reactive oxygen species (ROS) (PubMed:22268143). Early but transient accumulation after osmotic stress (e.g. polyethylene glycol, PEG) (PubMed:23815736). Induced by SA; early induction is NPR1-independent, but full-scale induction is NPR1-dependent (PubMed:14742872, PubMed:22325892, PubMed:22268143, PubMed:26433201). Up-regulated by benzothiadiazole (BTH) (PubMed:26433201). Repressed by jasmonic acid (MeJA) by both COI1-dependent and COI1-independent pathways (PubMed:14742872, PubMed:18713432). Triggered by the pathogenic compatible bacteria E.carotovora subsp. carotovora SCC3193 (PubMed:14742872). Induced by P.syringae pv. tomato DC3000 (PubMed:17965588, PubMed:22325892). Stimulated by ATX1 (PubMed:17965588). Up-regulated by E.amylovora (PubMed:22316300). Accumulates during leaf and flower senescence (PubMed:17310369). Induced expression upon simultaneous feeding by caterpillars (e.g. P.xylostella) and aphids (e.g. B.brassicae) at a low density, but lower levels in plants induced with both caterpillars and a high aphid density (PubMed:25339349). Responsive to rhizobacterium B.cereus AR156 in leaves (PubMed:26433201). {ECO:0000269|PubMed:14742872, ECO:0000269|PubMed:17310369, ECO:0000269|PubMed:17965588, ECO:0000269|PubMed:18713432, ECO:0000269|PubMed:22268143, ECO:0000269|PubMed:22316300, ECO:0000269|PubMed:22325892, ECO:0000269|PubMed:23067202, ECO:0000269|PubMed:23603962, ECO:0000269|PubMed:23815736, ECO:0000269|PubMed:25339349, ECO:0000269|PubMed:26433201}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002961829.2 | 7e-43 | probable WRKY transcription factor 31 | ||||
| Swissprot | Q9LY00 | 3e-25 | WRK70_ARATH; Probable WRKY transcription factor 70 | ||||
| TrEMBL | D8QTC4 | 6e-44 | D8QTC4_SELML; Uncharacterized protein WRKY_25 (Fragment) | ||||
| STRING | EFJ18006 | 1e-44 | (Selaginella moellendorffii) | ||||
| STRING | EFJ37089 | 1e-44 | (Selaginella moellendorffii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP14 | 17 | 875 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G56400.1 | 1e-27 | WRKY DNA-binding protein 70 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 29692 |
| Entrez Gene | 9629159 | 9657340 |



