 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00015267-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 477aa MW: 52808.2 Da PI: 7.2396 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 31.8 | 3.2e-10 | 173 | 202 | 4 | 33 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLee 33
k rr+++NReAAr+sR+RKka++++Le
SMil_00015267-RA_Salv 173 AKTLRRLAQNREAARKSRLRKKAYVQQLES 202
5889************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 6.0E-5 | 170 | 247 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 2.0E-7 | 172 | 216 | No hit | No description |
| PROSITE profile | PS50217 | 9.117 | 172 | 216 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 2.4E-7 | 173 | 208 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.93E-6 | 174 | 216 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 177 | 192 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 7.8E-33 | 255 | 330 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 477 aa Download sequence Send to blast |
MASHRVGEAD SGAPHHHHIP YAIPHMLHHP NNSTFINQEG SAFDFGELEE AIILQGVKIN 60 NDDKKQPLYT AIRPAATLDM FPSWPMRFQQ HTLRGSSKSG EESTDSGSAV NTLSNTPESP 120 SDHRQPPPPP PQMDAAGGID SPRLSGSHSQ PAAKPAPEKR RGPGSNSDKV LDAKTLRRLA 180 QNREAARKSR LRKKAYVQQL ESSRIRLAQL EQDLQRARSQ GLFLGGGGAA SGSISSGGAI 240 FDMEYSRWLD DDHRHMSELR TALQAHLSDG DLRVIVDGYI AHYDEVFRLK GAAAKSDVFH 300 LITGMWTTPA ERCFLWMGGF RPSDLIKMLI PQLDPLTEQQ FMGICSLQHS SQQAEEALSQ 360 GLEQLQQSLV DTIANGSAND SMHHMAVALG KLANLEGFVR QADNLRQQTL HQLHRLLTIR 420 QAARCFLVIG EYYGRLRALS SLWASRPRES MIADDNSCQT TTDLQMVQSS QNQFSNF |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Together with TGA10, basic leucine-zipper transcription factor required for anther development, probably via the activation of SPL expression in anthers and via the regulation of genes with functions in early and middle tapetal development (PubMed:20805327). Required for signaling responses to pathogen-associated molecular patterns (PAMPs) such as flg22 that involves chloroplastic reactive oxygen species (ROS) production and subsequent expression of H(2)O(2)-responsive genes (PubMed:27717447). {ECO:0000269|PubMed:20805327, ECO:0000269|PubMed:27717447}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00131 | DAP | Transfer from AT1G08320 | Download |
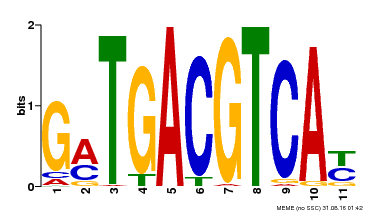
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by flg22 in leaves. {ECO:0000269|PubMed:27717447}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011088884.1 | 0.0 | transcription factor TGA9 isoform X2 | ||||
| Swissprot | Q93XM6 | 0.0 | TGA9_ARATH; Transcription factor TGA9 | ||||
| TrEMBL | A0A4D8ZWL6 | 0.0 | A0A4D8ZWL6_SALSN; Transcription factor TGA | ||||
| STRING | EOY33449 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3683 | 23 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08320.3 | 1e-172 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



