 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00009551-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 185aa MW: 20153.6 Da PI: 6.2664 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 61 | 2.7e-19 | 36 | 85 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
ykGVr++ +g+WvAeIr+p + r r++lg+f+t ++Aa a++aa++kl+g
SMil_00009551-RA_Salv 36 TYKGVRQRT-WGKWVAEIREP---N-RgSRVWLGTFDTSYDAAVAYDAAARKLYG 85
69****999.**********8...3.35*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.72E-18 | 35 | 95 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.0E-30 | 36 | 94 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.023 | 36 | 93 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 9.0E-13 | 36 | 85 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 9.6E-36 | 36 | 99 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.29E-21 | 36 | 94 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 1.3E-9 | 37 | 48 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-9 | 59 | 75 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 185 aa Download sequence Send to blast |
MLTAAGEEKR ARKIAQASSR KGCMRGKGGP ENAACTYKGV RQRTWGKWVA EIREPNRGSR 60 VWLGTFDTSY DAAVAYDAAA RKLYGPDAKL NLPHLYEEMP PHQPPQLPAE ATSGGYYPAP 120 ADCGEYVSGC GILKNLDVSL PEVDDSYLWA EAAKGTSFRI VEEEGAFATN LDGNGVPFPW 180 FPHSL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 6e-17 | 37 | 93 | 6 | 63 | ATERF1 |
| 3gcc_A | 6e-17 | 37 | 93 | 6 | 63 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Putative transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity-inducible transcription. {ECO:0000269|PubMed:11798174}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00240 | DAP | Transfer from AT1G75490 | Download |
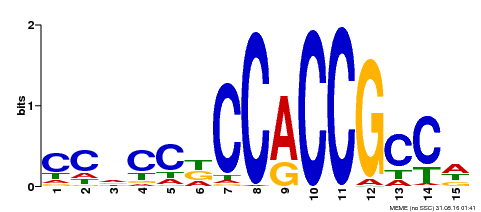
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By high-salt stress. {ECO:0000269|PubMed:11798174}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011083628.1 | 6e-88 | dehydration-responsive element-binding protein 2D-like | ||||
| Swissprot | Q9LQZ2 | 3e-51 | DRE2D_ARATH; Dehydration-responsive element-binding protein 2D | ||||
| TrEMBL | A0A4D9A0H4 | 8e-94 | A0A4D9A0H4_SALSN; Uncharacterized protein | ||||
| STRING | Migut.C00778.1.p | 5e-67 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3155 | 22 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75490.1 | 7e-47 | ERF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



