 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00008639-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 451aa MW: 49444.2 Da PI: 6.4177 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51 | 3.4e-16 | 78 | 122 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++++a k++G W++I +++g +t+ q++s+ qk+
SMil_00008639-RA_Salv 78 RERWTEEEHKKFLEALKLYGRA-WRRIEEHVG-SKTAVQIRSHAQKF 122
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.05E-16 | 72 | 127 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.057 | 73 | 127 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 4.1E-17 | 76 | 125 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 4.0E-13 | 77 | 125 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-13 | 78 | 121 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-9 | 78 | 118 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.86E-10 | 80 | 123 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 451 aa Download sequence Send to blast |
LFREEFGESD CFSAAMVIKG VIGGEECIVS GSPLPTNDEI SLDLDGQDVK DVQLNEQFAG 60 GDDFAPKVRK PYTITKQRER WTEEEHKKFL EALKLYGRAW RRIEEHVGSK TAVQIRSHAQ 120 KFFSKVARES NVGDGGLVKP VEIPPPRPKR KPMHPYPRKL GSPVKMGILI PERSASPNQS 180 PTSVLSVVGS DQSSGGADSC MPNGSSSPVS SSPLARTTAH IFGSEAPKLL SEDGVSSSQS 240 DREDENSSTD EEIPLELDLS PRKTFVNEAA TQSLKLFGKT LLVVDSSRST CNPESLDDDK 300 RVETCPYPLR VVPLKAAGNS TCEDAWRSLM QVHSSEMKNE LLYIQYTGNT TSSSAAPFPW 360 LSLASQSTRE VHSPTPIRAQ SNEGSSSGSN TDSVGAAARE ETSSFSSYKL SKRISANCRR 420 GFVPYKRCFT EQDSSAETVQ EREKQRIRLC L |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
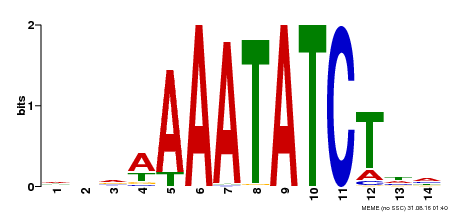
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011085926.1 | 1e-136 | protein REVEILLE 1 | ||||
| TrEMBL | A0A4D9B7Y1 | 0.0 | A0A4D9B7Y1_SALSN; MYB-related transcription factor LHY | ||||
| STRING | XP_009790907.1 | 1e-89 | (Nicotiana sylvestris) | ||||
| STRING | Migut.K00742.1.p | 6e-91 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3185 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 1e-57 | MYB_related family protein | ||||



