 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00003982-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 317aa MW: 33767 Da PI: 8.4186 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 107.1 | 2.7e-33 | 40 | 165 | 1 | 123 |
TCP 1 aagkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec..eaesssssas..n 84
a+g+kdrhsk+ T++g+RdRRvRls+++a++f+dLqd+LG+d++sk++eWLl +a +i el+ +s+ a+++ + + + + +
SMil_00003982-RA_Salv 40 ATGGKDRHSKVLTSRGLRDRRVRLSVNTAIEFYDLQDRLGYDQPSKAVEWLLAAAAGSIAELPPMNSPFAESAaaS--GRMMALDsvF 125
5899*************************************************************77777444221..2222111122 PP
TCP 85 sssg....kaaksaakskksqksaasalnlakesrakararar 123
+ +a ks+ s++s++s+ s l++a++ +ak+r++++
SMil_00003982-RA_Salv 126 ---DggeaAAGKSSIGSSSSETSKGSGLSIARSDKAKERENEK 165
...1343355555558888889999889999999988888754 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.6E-32 | 42 | 160 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 30.555 | 43 | 101 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MNDDDEGENR RSNPNKGGSA DPVGGLYGWP SSRIVRVSRA TGGKDRHSKV LTSRGLRDRR 60 VRLSVNTAIE FYDLQDRLGY DQPSKAVEWL LAAAAGSIAE LPPMNSPFAE SAAASGRMMA 120 LDSVFDGGEA AAGKSSIGSS SSETSKGSGL SIARSDKAKE RENEKSSYVA SFTELLSGGG 180 GGINCFNGRD QRPVNEENSN FFLKFPTDYF SRGLSAPQAT ASPSPLLFSA AAELQPPFPF 240 ASDQSVSAVN AANPSAINRG TLQSNSSSPP HFPPNFFGSG NHRHFFPGFD SRLQLYYGDA 300 DQGNDGRNSG PKRKDKP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 2e-21 | 48 | 102 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 2e-21 | 48 | 102 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00169 | DAP | Transfer from AT1G30210 | Download |
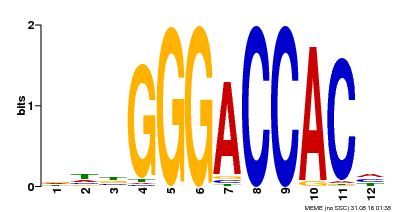
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012830233.1 | 7e-77 | PREDICTED: transcription factor TCP2-like | ||||
| Refseq | XP_012842244.1 | 7e-76 | PREDICTED: transcription factor TCP2-like isoform X1 | ||||
| TrEMBL | A0A4D9AS47 | 2e-81 | A0A4D9AS47_SALSN; Uncharacterized protein | ||||
| STRING | Migut.H01353.1.p | 3e-76 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2464 | 22 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G30210.2 | 7e-40 | TCP family protein | ||||



