 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00003681-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 596aa MW: 65053.8 Da PI: 6.2549 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 68.8 | 8.4e-22 | 462 | 524 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
++elk e+rkq+NRe+ArrsR+RK+ae eeL++kv++L +eN++Lk+el+e k ++kl+ e+
SMil_00003681-RA_Salv 462 QPELKWEKRKQSNRESARRSRLRKQAEAEELATKVQTLITENMTLKSELDEFMKFSKKLELEN 524
5799****************************************************9998876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14223 | 5.6E-19 | 28 | 140 | No hit | No description |
| Pfam | PF07777 | 2.8E-19 | 206 | 284 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Gene3D | G3DSA:1.20.5.170 | 2.7E-13 | 460 | 520 | No hit | No description |
| SMART | SM00338 | 1.1E-18 | 462 | 526 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 2.5E-19 | 463 | 524 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.243 | 464 | 527 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.94E-9 | 466 | 518 | No hit | No description |
| CDD | cd14702 | 4.80E-18 | 468 | 515 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 469 | 484 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 596 aa Download sequence Send to blast |
MHSVKEKVNK ALLTKIDKTV TDEKLHDMND NARATILLNL SSSVVRKVSH HACAKELWDE 60 LNSIYAAPSE VSTWSLQNQF MSFQMDSSKD VDTNMEIFNK LLHDLKLAGD DSIEKYAPQI 120 LLNSIPESFV EFISYSNAIS KTMSSSEWLI DSGCTFHVTS FEHMIHKLQP VKSGHVALAD 180 GQNCEIIGKA ARWSSEIPEE LRRIRQSVAE QDQSAAPVYH DWAMMQAYYG PQFAVPPYLN 240 SAVASPHIPT PYMWMVPPQY TVPPYGASYP GYYSHEGAYG HPGVSTAGTS FSMATPAESS 300 GNTDGGIVKK KSKDSQGLAV PIENGNADGG GHGPDRRLSE SEVTDDSSDG TNEVTAEAGQ 360 SGVKRSLAGS PKSGKLLLSS SLSISAAKGD KDQKKGSADV GRGYGSTEEA SRPAYTPAKA 420 KENASTVPEL KGSSILDVKP VAACNPRSSI ANEAQLQGTP NQPELKWEKR KQSNRESARR 480 SRLRKQAEAE ELATKVQTLI TENMTLKSEL DEFMKFSKKL ELENGTLMEK LKDARPGEVD 540 FLKIDGVRPK PVDTVNLLAR VESNGATERR DEDGDSYEQR SRAAKLHQLL DAVAAG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 478 | 484 | RRSRLRK |
| 2 | 478 | 485 | RRSRLRKQ |
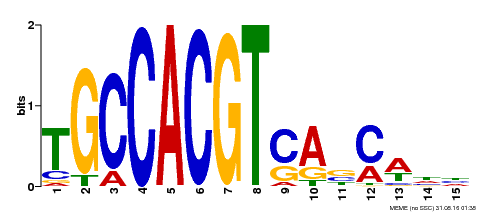
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020551515.1 | 1e-121 | common plant regulatory factor 1 isoform X2 | ||||
| TrEMBL | A0A4D9AV77 | 0.0 | A0A4D9AV77_SALSN; Plant G-box-binding factor | ||||
| STRING | Migut.B01828.1.p | 1e-102 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3113 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.2 | 8e-28 | G-box binding factor 3 | ||||



