 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sme2.5_07791.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 526aa MW: 58529.3 Da PI: 8.0079 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 53.9 | 3.1e-17 | 333 | 379 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+ellP++ +K +Ka++L +A+eY+ksLq
Sme2.5_07791.1_g00001.1 333 VHNLSERRRRDRINEKMKALQELLPHS-----TKTDKASMLDEAIEYLKSLQ 379
6*************************9.....9******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 18.476 | 329 | 378 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.09E-18 | 332 | 383 | No hit | No description |
| SuperFamily | SSF47459 | 3.53E-20 | 332 | 406 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.3E-14 | 333 | 379 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.7E-20 | 333 | 387 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.0E-18 | 335 | 384 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 526 aa Download sequence Send to blast |
MNPCLPEWNM ETELPVPHQK KSMGFDHELV ELLWRNGEVV LHSQTHKKHP GYDPNECRQF 60 NKHDQPTLRV AGNQTNFIRD DETVSWLNCP IDDSSFDKEF CSPFLSDIST TPPLGEAAHK 120 SVRHTEDNNK VYKFDPSGIN HVLPQAHHSG FDPNPMPPPR FHNFGSVQQQ QHHTVGQPQK 180 GVNFPPPVRS SNVQLSGKEA RSNLMKEGSV RTVGSSHCGS NQVDTSRFSS SANRGLSAAM 240 ITDYTGKISP QSDTMDRDTF EPANTSSSSG RSGSSYARAC NQSTATNSQS HKRKSRDGEE 300 PECQSKADEL ESAGGNKSAQ KSGTARRSRA AEVHNLSERR RRDRINEKMK ALQELLPHST 360 KTDKASMLDE AIEYLKSLQM QLQMMWMGSG MASMMFPGVQ HYMSRMGMGM GPPSVPSIHN 420 ALHLTRLPLV DPSIPLTQAA PNNQAAMCQN SMLNQVNYQC HLQNPNFPDQ YASYMGFHPL 480 QGASQPAMNI FGLGSHTAQQ SQQLPNPTNR QGNCFIWPIV YHGTRL |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 337 | 342 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
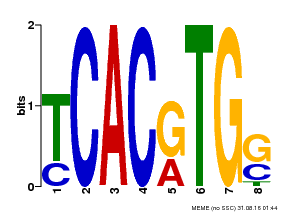
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013582 | 0.0 | BT013582.1 Lycopersicon esculentum clone 132330F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001294937.1 | 0.0 | transcription factor PIF4 | ||||
| Refseq | XP_015082675.1 | 0.0 | transcription factor PIF4 | ||||
| Refseq | XP_019070178.1 | 0.0 | transcription factor PIF4 isoform X1 | ||||
| Refseq | XP_027774395.1 | 0.0 | transcription factor PIF4 | ||||
| TrEMBL | A0A3Q7H9E8 | 0.0 | A0A3Q7H9E8_SOLLC; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400041156 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 7e-38 | phytochrome interacting factor 4 | ||||



