 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sme2.5_06702.1_g00005.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 379aa MW: 41562.8 Da PI: 7.3443 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.6 | 2.5e-32 | 198 | 253 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs+ AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Sme2.5_06702.1_g00005.1 198 KQRRCWSPELHRRFLHALQQLGGSHAATPKQIRELMKVDGLTNDEVKSHLQKYRLH 253
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.542 | 195 | 255 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.4E-18 | 196 | 256 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-29 | 196 | 256 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.8E-27 | 198 | 253 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.5E-7 | 200 | 251 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 379 aa Download sequence Send to blast |
MERCQQYIDA LEEERKKIQV FSRELPLCLE LVIQAIETYK QQLSGTTTEY NLNAQSTECS 60 EDEHTSSDVP IFEEIIPLKR TFAHEDEEDE SHKSKSFNNG NNNSNPKKSD WLRSVQLWNQ 120 TSDPTPKEEL TSEKVSVVEV KKHGSGGAFH PFKKEKSTVA LAKTPDTSGI VPAATGSSTA 180 GNNSGGENKK EDKDGQRKQR RCWSPELHRR FLHALQQLGG SHAATPKQIR ELMKVDGLTN 240 DEVKSHLQKY RLHTRRPSPS SIHNNNNQQP PQFVVVGGIW VPPPDYAGMA AAAPAASGDA 300 SGVANSNGIY APIATHPKGP LHDHALGGTL KQLRQHNKKS SRSSDERDDS HCHSDGGGVH 360 SNSPATSSST HTTTASPAY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 4e-13 | 198 | 251 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-13 | 198 | 251 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-13 | 198 | 251 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-13 | 198 | 251 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-13 | 198 | 251 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
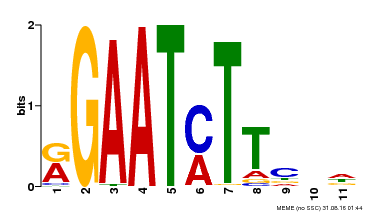
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015074811.1 | 0.0 | transcription factor HHO2 | ||||
| Swissprot | Q9FPE8 | 8e-98 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A1U8EQX2 | 0.0 | A0A1U8EQX2_CAPAN; transcription factor LUX | ||||
| STRING | Solyc05g009720.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA5219 | 24 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 3e-75 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



