 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Solyc08g060840.1.1 | ||||||||
| Common Name | LOC101258392 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum; Lycopersicon
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 827aa MW: 94634.5 Da PI: 7.5858 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 87.7 | 1.7e-27 | 67 | 152 | 2 | 90 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHela 90
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g++ e+++ ++r+ ++++t+Cka+++vk+++dgkw+v+++ ++HnH l
Solyc08g060840.1.1 67 FYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGTTPESDTG---SSRRPSVKKTDCKASMHVKRKRDGKWYVHEFIKDHNHGLL 152
9***************************************999877...77888899*****************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 5.2E-25 | 67 | 153 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 6.5E-25 | 273 | 365 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 10.039 | 553 | 589 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 8.1E-7 | 563 | 588 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.2E-8 | 564 | 591 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 827 aa Download sequence Send to blast |
MVDHGDVVQS SVQLTGDMVD AVDKSCLSRD GGVSRSPKRS ITGVEEHADF EPHDGIEFES 60 HEAAYAFYQE YAKSMGFTTS IKNSRRSKKS KEFIDAKFAC SRYGTTPESD TGSSRRPSVK 120 KTDCKASMHV KRKRDGKWYV HEFIKDHNHG LLPALAYHFR IHRNVKLAEK NNIDILNAVS 180 ERTRKMYVEM SRQCGGSQEV GLLTNDLNYQ FDKGRCLSLE EGDAHIMLEY FMHVQKENPC 240 FFYATDLNED QRLRNLFWID AKSRKDYVSF NDVVFFDTSY MKSNEKMPFA LLIGVNHHCQ 300 PMLLGCALIA DETKPTFVWL MKTWLRAVGG KAPKVIIADQ DKSLKSALEE VFPCSSHCFA 360 LWHVLERIPE TLAHVVKQHE NFMQKFSKCI FKSLTDEQFD LRWWKMVSRF ELQENEWIHT 420 LYEDRKKWIP AYMRGSFMAG MSTAQRSESI SSFFDKYIHK KISLKEFMRQ YGMILQNRYE 480 EEAIADFDTL HKLPALKSPS PWEKQMSTIY THTIFKKFQV EVLGVVGCHP KKEAVNGENV 540 TFRVDDCEKD ENFMVTWNEA RSDVSCSCLL FEYNGFLCRH AMIVLQMCGL SIIPSQYILK 600 RWTKDAKNIQ LISEGTERIR NRVQRYNDLC RRAIELGVEG SLSEESYGVA FRALDEALKN 660 CVNVNNRSSA LTECSSSAVG LRDLEEDTQG IHAIKTSRKK NTNKKRKMHS EPEAAIVEAK 720 DSLQQMDSLT VGGMTLNGYY GTHQNVQGLI QLNLMEPPHD GYYVNQQNMQ GLGQLNTIAP 780 GHDGFFGSQQ SIPGLGHLDF RQPSFTYGLQ DEPSLRAAQL HGNNAR* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Les.8437 | 0.0 | callus| flower | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
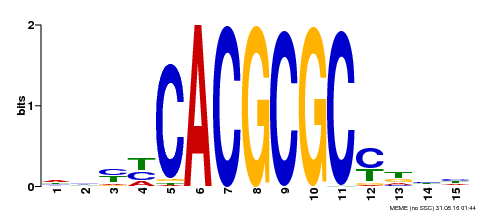
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Solyc08g060840.1.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004245025.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Refseq | XP_010324981.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Refseq | XP_010324983.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | K4CKM9 | 0.0 | K4CKM9_SOLLC; Uncharacterized protein | ||||
| STRING | Solyc08g060840.1.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 | Representative plant | OGRP13457 | 3 | 4 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Solyc08g060840.1.1 |
| Entrez Gene | 101258392 |



