 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Solyc01g108080.2.1 | ||||||||
| Common Name | LOC101263766 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum; Lycopersicon
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 415aa MW: 45028.3 Da PI: 10.2455 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 41 | 4e-13 | 336 | 391 | 5 | 59 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH.HHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke.leelkkevakl 59
+r rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+k+ +e ++k+ +l
Solyc01g108080.2.1 336 RRRRRMIKNRESAARSRARKQAYTLELEAEVAKLKEINEELRKKqAEIIEKQKNQL 391
689**************************************976134445555555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.3E-11 | 332 | 393 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.841 | 334 | 383 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.5E-12 | 336 | 386 | No hit | No description |
| SuperFamily | SSF57959 | 2.81E-9 | 336 | 383 | No hit | No description |
| Pfam | PF00170 | 5.0E-11 | 336 | 391 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 339 | 354 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 2.83E-22 | 341 | 390 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 415 aa Download sequence Send to blast |
MGSYMNFKNI TDKPQAESNG GKSVGNGDIP LARQSSIYSL TFDELQTTFS GLGKDFGSIN 60 MEELLKSIWT AEESQAATSS TGGGEDGIAP VGNLQRQGSL TLPRTLSQKT VDEVWRNFQK 120 ETTVCTPDGS ETGKSNFGQR QSTLGEMTLE EFLVKAGVVR EDMQSTSNSS GITFNNGLSQ 180 QNNNNGFNIA FQQPTQNNGL LINQIAANNM LNVVGATASQ QQQPQQQQPL FPKQTTVAFA 240 SPMQLSNNGH LASPRTRAPA VGMSSPSVNA SMAQGGVMGK TGFHNGVSPA KVGSPGNDFI 300 ARSNVDTSSL SPSPYAFSEG GRGRRSGSSL EKVVERRRRR MIKNRESAAR SRARKQAYTL 360 ELEAEVAKLK EINEELRKKQ AEIIEKQKNQ LTDKRNMTCG YKLRCLRRTL TGPW* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, leaves, flowers and siliques but not in seeds. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
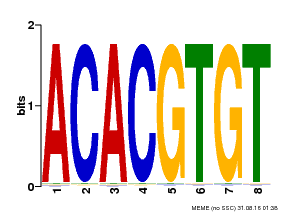
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Solyc01g108080.2.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA), cold and glucose. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004230778.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Swissprot | Q9M7Q4 | 4e-99 | AI5L5_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | V5L071 | 0.0 | V5L071_SOLNI; Abscisic acid-insensitive 5-like protein | ||||
| STRING | Solyc01g108080.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA776 | 24 | 97 | Representative plant | OGRP3984 | 10 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 4e-81 | abscisic acid responsive elements-binding factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Solyc01g108080.2.1 |
| Entrez Gene | 101263766 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



