 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.9G524200.1.p | ||||||||
| Common Name | LOC101768133, SETIT_036519mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 336aa MW: 36880 Da PI: 6.5483 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.7 | 8.2e-18 | 113 | 166 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rqV +WFqNrRa++k
Seita.9G524200.1.p 113 KKRRLNVEQVRTLEKNFELGNKLEPERKLQLARALGLQPRQVAIWFQNRRARWK 166
4568999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 127.9 | 4.2e-41 | 112 | 202 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
ekkrrl+ eqv++LE++Fe +kLeperK +lar+Lglqprqva+WFqnrRAR+ktkqlEkdy+aLkr++da+k++n++L +++++L++e+
Seita.9G524200.1.p 112 EKKRRLNVEQVRTLEKNFELGNKLEPERKLQLARALGLQPRQVAIWFQNRRARWKTKQLEKDYDALKRQLDAVKADNDALLSHNKKLQAEI 202
69**************************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-18 | 91 | 169 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.92E-19 | 104 | 170 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.346 | 108 | 168 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.3E-17 | 111 | 172 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.46E-16 | 113 | 169 | No hit | No description |
| Pfam | PF00046 | 3.3E-15 | 113 | 166 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 1.6E-5 | 139 | 148 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 143 | 166 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.6E-5 | 148 | 164 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.2E-14 | 168 | 208 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MRPMASNGMA SSPSPFFPPN FLLQMQQTPH DHDPQEHHHH HHEHHLPPPL HPHHNPFLSS 60 SQCPSLQDFR GMAPMLGKRP MYGADVGGGD ETNGGGGVNE DELSDDGSQA GEKKRRLNVE 120 QVRTLEKNFE LGNKLEPERK LQLARALGLQ PRQVAIWFQN RRARWKTKQL EKDYDALKRQ 180 LDAVKADNDA LLSHNKKLQA EILALKGREA GSELINLNKE TEASCSNRSE NSSEINLDIS 240 RTPPSEGPMD PPPPDHQHPS SGGGGGGGMI PFYPSAGRPS GVDIDQLLHT TSVPKLEQHS 300 GGGVQGAETA SFGNLLCGVD EPPPFWPWAD HQHFH* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 160 | 168 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
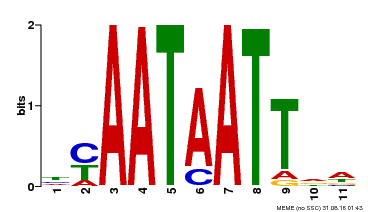
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.9G524200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU966490 | 0.0 | EU966490.1 Zea mays clone 294849 homeobox-leucine zipper protein HAT7 mRNA, complete cds. | |||
| GenBank | KJ728506 | 0.0 | KJ728506.1 Zea mays clone pUT6811 HB transcription factor (HB102) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004985543.1 | 0.0 | homeobox-leucine zipper protein HOX21 isoform X2 | ||||
| Swissprot | Q8S7W9 | 1e-148 | HOX21_ORYSJ; Homeobox-leucine zipper protein HOX21 | ||||
| TrEMBL | K4ACB2 | 0.0 | K4ACB2_SETIT; Uncharacterized protein | ||||
| STRING | Si036519m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2289 | 38 | 94 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-77 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.9G524200.1.p |
| Entrez Gene | 101768133 |



