 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.8G135700.4.p | ||||||||
| Common Name | LOC101781693, SETIT_025988mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 812aa MW: 90033.3 Da PI: 7.052 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.1 | 3.4e-23 | 112 | 213 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d++g +W++++i+r++++r++l++GW+ Fv+a++L +gD ++F
Seita.8G135700.4.p 112 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHGAEWRFRHIFRGQPRRHLLQSGWSVFVSAKRLVAGDAFIFL-- 199
99*****************************7333.34459************************************************.. PP
SSSEE..EEEEE-S CS
B3 86 grsefelvvkvfrk 99
+ ++el+v+v+r+
Seita.8G135700.4.p 200 RGVNGELRVGVRRA 213
4489999****996 PP
| |||||||
| 2 | Auxin_resp | 105.8 | 5.1e-35 | 238 | 320 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++t+++F+v+Y+Pr+s++eFvv++++++++lk+++ +GmRfkm+fe+e+++e+r++Gt+vg d d+ Wp+SkWr Lk
Seita.8G135700.4.p 238 AWHAVNTGTMFTVYYKPRTSPAEFVVPCDRYMESLKRNYPIGMRFKMRFEGEEAPEQRFTGTIVGNVDPDQAGWPESKWRYLK 320
89******************************************************************************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 8.8E-42 | 104 | 227 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.67E-40 | 105 | 241 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.68E-21 | 110 | 212 | No hit | No description |
| SMART | SM01019 | 2.6E-23 | 112 | 214 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 3.6E-21 | 112 | 213 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.519 | 112 | 214 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.4E-32 | 238 | 320 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 21.208 | 686 | 768 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 2.89E-11 | 687 | 762 | No hit | No description |
| Pfam | PF02309 | 7.6E-8 | 727 | 771 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
MYTELWNLCA GPLVTVPSVG DKVYYFPQGH IEQVEASTNQ VAEQHMQLYN LPWKILCEVM 60 NVELKAEPDT DEVYAQLTLL PESKQPDENG SSEQEMPAAP SAAPARPRVH SFCKTLTASD 120 TSTHGGFSVL RRHADECLPP LDMSRQPPTQ ELVAKDLHGA EWRFRHIFRG QPRRHLLQSG 180 WSVFVSAKRL VAGDAFIFLR GVNGELRVGV RRAMRQQANV PSSVISSHSM HLGVLATAWH 240 AVNTGTMFTV YYKPRTSPAE FVVPCDRYME SLKRNYPIGM RFKMRFEGEE APEQRFTGTI 300 VGNVDPDQAG WPESKWRYLK VRWDEASSIP RPERVSPWQI EPAVSPPPIN PLPVHRPKRP 360 RSNAVASLPD SSAPTKEAAP KVTVEAQQNA LQRVLQTQDN ATPKSAFGDK SELDAAQQSV 420 LRPSGFDREK STIGTQRKLG SDSWMQMSRP ESYNEMLSGL SGYQQPKDLQ NQQGFCSLPD 480 QIAAGRPNFW HTVNAHYQDQ QGNHNMFGSW SMMPSSTGFG LNRQNYPMIQ EVGAMPQSSA 540 NTKFGNGVYT PLPGRGIDQY SAGWFGHMVP GSRMDDAQPR VIKPQPLVLA HGDAQKMKGN 600 SCKLFGIHLD SPAKSEPLKS PPSVANDGMP QTPAAAEWRR VDTTEVEKSS DPPKTPKQLD 660 APQADPVPCP QSSRSTQCKS QGGSTRSCKK VHKQGIALGR SVDLTKFKGY TELVSELDEM 720 FDFNGELKGS NKEWMVVYTD NEGDMMLVGD DPWDEFCNMV HKIFIYTTEE VQRMNPGTLN 780 SGSEDSPANS MERGSAVRET LPASSLNSGN C* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-170 | 4 | 342 | 22 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-170 | 4 | 342 | 22 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-170 | 4 | 342 | 22 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-170 | 4 | 342 | 22 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-170 | 4 | 342 | 22 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
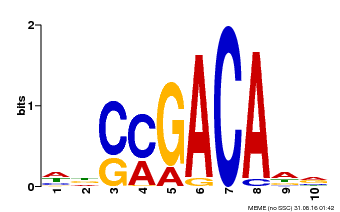
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.8G135700.4.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022684732.1 | 0.0 | auxin response factor 23 isoform X1 | ||||
| Refseq | XP_022684733.1 | 0.0 | auxin response factor 23 isoform X1 | ||||
| Swissprot | A2ZET6 | 0.0 | ARFW_ORYSI; Auxin response factor 23 | ||||
| TrEMBL | K3ZHD6 | 0.0 | K3ZHD6_SETIT; Auxin response factor | ||||
| STRING | Si025988m | 0.0 | (Setaria italica) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.8G135700.4.p |
| Entrez Gene | 101781693 |



